The suitability of small biopsy and cytology specimens for EGFR and other mutation testing in non-small cell lung cancer
Introduction
Lung cancer is the commonest cause of cancer death worldwide, being responsible for an estimated 1.59 million deaths in 2012 (1). Non-small cell lung cancer (NSCLC) comprises the majority of lung cancer cases, and around 70% of patients present at an advanced stage, when surgical resection is no longer a treatment option (2).
Traditionally, platinum-based chemotherapies were the mainstay of treatment for advanced-stage NSCLC. The past decade however has seen a major paradigm shift in the treatment approach for advanced NSCLC. In 2004, reports emerged of activating somatic mutations in the epidermal growth factor receptor (EGFR) gene in NSCLC, conferring sensitivity to treatment with EGFR tyrosine kinase inhibitors (TKIs) (3-5). Subsequent phase III trials confirmed improved response rates and progression-free survival in NSCLC patients treated with EGFR TKIs compared to platinum-based chemotherapies, where the tumor harbored an activating EGFR mutation (6,7).
EGFR is a transmembrane receptor tyrosine kinase, which activates downstream pathways involved in cellular proliferation and survival. Activating mutations in the tyrosine kinase domain, the majority of which involve exon 19 deletions or a point mutation (L858R) in exon 21, result in enhanced sensitivity to EGFR TKIs, which occupy the ATP binding site (3). Mutations conferring resistance to EGFR TKIs are also reported, notably exon 20 insertions and the exon 20 T790M point mutation (8-11).
Mutation testing has emerged as a vital element in the diagnostic work-up of advanced NSCLC patients, in order to determine which patients are likely to benefit from treatment with targeted therapies. Direct sequencing has been considered the reference method, but is limited by low analytical sensitivity compared to newer platforms (12), which is a particular problem with the typically low cellularity of many lung cancer biopsies. Our institution utilizes a commercially available multiplex PCR assay with analysis based on matrix-assisted laser desorption ionization-time of flight (MALDI-TOF) mass spectrometry technology (13,14).
Although the primary focus is to detect EGFR mutations, our institution also reports mutations in KRAS and BRAF detected by this multiplex method, on the basis that they may be of clinical relevance in NSCLC. Clinical trials of BRAF inhibitors in NSCLC are currently in progress (11). Although no targeted therapies are available for KRAS mutations in NSCLC (11), the presence of a KRAS mutation effectively excludes an EGFR mutation and negates the need to pursue ALK testing, as such major driver mutations are almost always mutually exclusive (9,10).
In the first instance, NSCLC is typically diagnosed on a small biopsy or cytology specimen, obtained through a minimally invasive procedure such as a CT-guided biopsy or bronchoscopy. As most patients with NSCLC present with advanced-stage disease not suitable for surgical resection, these small biopsy or cytology specimens are often the only samples available for mutation testing. The utility of these different specimens in comparison to resected tumors (the “gold standard”) remains to be fully elucidated.
This study reviews the NSCLC cases submitted to our institution for somatic mutation testing, and compares the mutation profiles of different specimens with the aim of assessing their suitability for mutation testing in a real world setting.
Materials and methods
Patients
This study was approved by the Human Research Ethics Committee of Royal Prince Alfred Hospital. Mutation testing undertaken on NSCLC cases between March 2012 and May 2013 at Royal Prince Alfred Hospital were retrospectively reviewed. All cases were referred by the treating physician. The supplied clinical and pathological information was recorded.
DNA extraction
DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) tissue, including cell blocks, and from a cell suspension in one case. Appropriate tumor blocks were selected by a pathologist and macrodissection was undertaken to increase the proportion of tumor cells where appropriate. The percentage of tumor cells in the tissue selected for mutation testing was estimated by an experienced pathologist (Sandra A. O’Toole or Wendy A. Cooper). DNA extraction was undertaken using NucleoSpin FFPE DNA Kit (Macherey Nagel, Düren, Germany). DNA quantity and quality was assessed using Nanodrop ND-1000 Spectrophotometer (NanoDrop Technologies, Wilmington, Delaware, USA) or Qubit 2.0 Fluorometer (Life Technologies, Carlsbad, California, USA). Following the manufacturer’s instructions, specimens with at least 500 ng of extracted DNA were deemed adequate for mutation testing (see below). Specimens with less than 500 ng of extracted DNA were either not tested further or tested with a qualifying comment about the risk of false negative or positive results, at the discretion of the supervising pathologists (Bing Yu, Sandra A. O’Toole, and Wendy A. Cooper).
Mutation testing
Samples were amplified using a 24-well multiplex PCR assay, OncoCarta Panel v1.0 Kit (Agena Bioscience, San Diego, California, USA), to assess mutations in EGFR, KRAS, and BRAF, with a sensitivity of 10%. PCR products were analyzed on a MALDI-TOF mass spectrometry-based platform, MassARRAY (Agena Bioscience, San Diego, California, USA). Fragment analysis of EGFR exons 19 and 20 was also undertaken to detect insertions or deletions in these regions. For fragment analysis, EGFR exons 19 and 20 were amplified by PCR with one fluorescent labeled primer and allele size was analyzed by capillary electrophoresis, using GeneMapper v4.0 (Life Technologies, Carlsbad, California, USA).
Statistical analysis
Specimen types were compared using an unpaired t-test (age and mean percentage of tumor cells), or Fisher’s exact test (gender, primary and metastatic tumors, adenocarcinoma histology, and mutation positive cases). Statistical analyses were performed using GraphPad Prism 6 (GraphPad Software, La Jolla, California, USA). A two-tailed P value of less than 0.05 was considered statistically significant.
Results
Clinicopathological characteristics
A total of 146 specimens were submitted for mutation testing, from 143 patients (three patients had two samples tested each). The clinicopathological characteristics of the patients are summarized in Table 1.
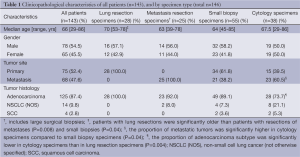
Full table
Specimen types
Small biopsy specimens comprised the highest proportion of cases (55 of 146; 37.7%), closely followed by resection specimens (53 of 146; 36.3%) and cytology specimens (38 of 146; 26%). Of the resection specimens, 28 were primary lung resections and 25 were resections (including large surgical biopsies) of a metastasis, including 13 brain, 6 pleura, and 3 lymph node specimens, and 1 specimen each of bone, mediastinum, and submandibular tissue. The small biopsy specimens included 50 core needle biopsies (including 29 lung, 8 lymph node, 7 bone, and 4 liver specimens, and 1 specimen each of abdominal wall and paravertebral soft tissue) and 5 bronchial biopsies. The cytology specimens included 14 percutaneous fine needle aspirations (FNAs) (11 lungs and 1 specimen each of adrenal gland, kidney, and lymph node), 8 endobronchial ultrasound-guided FNAs, 12 pleural and 1 pericardial fluid, and 3 bronchial washings.
Comparison of clinicopathological characteristics between specimen types
Age, gender, and tumor histology were compared between all specimen types and primary and metastatic site were compared between small biopsy and cytology specimens. Patients with lung resections were significantly older than patients with resections (including large surgical biopsies) of metastases (median age, 70 vs. 63; P=0.008), and patients with small biopsies (median age, 70 vs. 64; P=0.04). The proportion of adenocarcinoma subtype was significantly lower in cytology specimens than in lung resection specimens (73.7% vs. 100%; P=0.004). Cytology specimens comprised significantly more metastatic tumors compared to small biopsy specimens (60.5% vs. 38.2%; P=0.04). All other comparisons were not statistically significant.
Results of mutation testing
Of 146 specimens, 4 specimens (2.7%; 2 small biopsy and 2 cytology specimens) contained insufficient DNA for mutation testing and these specimens were excluded from subsequent analyses of mutations. There was no significant difference in the proportion of cases deemed insufficient between the specimen types.
The results of mutation testing are summarized in Table 2. Of the 142 evaluable specimens, EGFR mutations were detected in 31 specimens (21.8%), KRAS mutations in 31 specimens (21.8%) and BRAF mutations in 3 specimens (2.1%). EGFR, KRAS, and BRAF mutations were mutually exclusive.
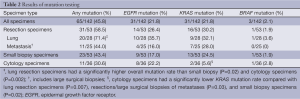
Full table
Resection specimens had the highest mutation rate (31 of 53 cases; 58.5%), including 14 cases (26.4%) with EGFR mutations, 16 cases (30.2%) with KRAS mutations and 1 case (1.9%) with BRAF mutation. The majority of mutations in resection specimens were found in lung resection specimens, of which 20 of 28 specimens (71.4%) harbored a mutation, including 10 cases (35.7%) with EGFR mutations, 9 cases (32.1%) with KRAS mutations, and 1 case (3.6%) with BRAF mutation.
Of 53 small biopsy specimens tested, mutations were detected in 23 cases (43.4%), including 9 cases (17%) with EGFR mutations, 13 cases (24.5%) with KRAS mutations, and 1 case (1.9%) with BRAF mutation. Mutations were detected in 11 of 36 (30.6%) cytology specimens, including 8 cases (22.2%) with EGFR mutations, 2 cases (5.6%) with KRAS mutations, and 1 case (2.8%) with BRAF mutation.
Three patients had two specimens tested each. In one case, an initial cytology specimen (FNA of the lung tumor), and a subsequent lung resection specimen were tested; both confirmed an EGFR L858R mutation. In another case, two small biopsy specimens were tested due to concerns about low DNA quantity in the first specimen; both specimens were negative. Finally in the third case, two resected synchronous primary tumors were tested; both contained a KRAS mutation (G12C in one tumor and G12D in the other).
Comparison of mutation rates
The overall mutation rate was significantly higher in lung resection specimens compared to small biopsy (71.4% vs. 43.4%; P=0.02), and cytology specimens (71.4% vs. 30.6%; P=0.002). However, there was no significant difference in the EGFR mutation rate between lung resection and small biopsy (35.7% vs. 17%; P=0.1) or lung resection and cytology specimens (35.7% vs. 22.2%; P=0.27).
There was no significant difference between small biopsy and cytology specimens, whether comparing the overall mutation rate (43.4% vs. 30.6%; P=0.27) or the EGFR mutation rate (17% vs. 22.2%; P=0.59).
The KRAS mutation rate was significantly lower in cytology specimens compared to lung resections (5.6% vs. 32.1%; P=0.007), resections (including large surgical biopsies) of metastases (5.6% vs. 28%, P=0.03), and small biopsy specimens (5.6% vs. 24.5%; P=0.02).
There was no significant difference between resections of lung and metastatic sites in comparing the overall (71.4% vs. 44%; P=0.055), EGFR (35.7% vs. 16%; P=0.13), or KRAS mutation rates (32.1% vs. 28%; P=0.77).
EGFR mutant cases
Of the 30 patients (31 specimens including one patient with two specimens) with EGFR mutations, there were 15 cases (50%) with isolated exon 21 L858R mutation, 9 cases (30%) with exon 19 deletions, 3 cases (10%) with exon 18 point mutation, 2 cases (6.7%) with both exon 21 L858R mutation and exon 20 T790M mutation, and 1 case (3.3%) with an exon 20 insertion. The EGFR mutation positive cases included 19 females (63.3%) and 11 males (36.7%), with a median age of 66.5 (range, 40-83), 28 cases (93.3%) of adenocarcinoma subtype and 2 cases (6.7%) of NSCLC (not otherwise specified), with 18 primary (60%) and 12 (40%) metastatic lesions.
Comparison of tumor cell content
Data on tumor cell content was available for 143 specimens (1 cell suspension and 2 specimens with unrecorded data were excluded). The mean percentage of tumor cells was highest in lung resection specimens (59.8%±3.4%; mean ± standard error of the mean), followed by resections (including large surgical biopsies) of metastases (54.6%±4.6%), cytology (40.4%±5.7%), and small biopsy specimens (36.7%±3.0%). There was a significantly higher mean percentage of tumor cells in lung resection specimens compared to small biopsy (P<0.0001) and cytology specimens (P=0.009), while there was no significant difference between small biopsy and cytology specimens (P=0.53).
There was no significant difference in the mean percentage of tumor cells among all specimens harboring EGFR mutations compared to those found to be EGFR wild type (53.3%±4.6% vs. 44.2%±2.5%; P=0.1). For cytology specimens alone (but not lung resections, metastatic resections, or small biopsy specimens alone), the mean percentage of tumor cells was significantly higher in cases harboring an EGFR mutation compared to those where no EGFR mutation was found (67.1%±12.6% vs. 35.5%±6.2%, P=0.03).
Discussion
This study retrospectively reviewed NSCLC mutation results with the types of specimens submitted for mutation testing at our institution. We compared the mutation rates between specimen types, to better define the suitability of small biopsy and cytology specimens for EGFR mutation testing. The majority (63.7%) of the submitted specimens were small biopsy or cytology specimens, reflecting the frequent use of such specimens for mutation testing (12). Indeed, a similar proportion of NSCLC patients are known to present with advanced-stage disease (2), and with a surgical resection specimen not forthcoming, mutation testing needs to be reliably performed on small biopsy and cytology specimens.
The overall mutation rates in our study (21.8% EGFR mutation, 21.8% KRAS mutation, 2.1% BRAF mutation) are consistent with published mutation rates (10,11), and reflect the predominantly Caucasian population of Sydney, Australia. The distribution of EGFR mutations, with the majority exon 19 deletions or exon 21 L858R mutations, is also consistent with known mutation rates (8-11).
Interestingly, lung resection specimens demonstrated a higher EGFR mutation rate of 35.7%. Presumably, a proportion of these cases represent patients with early-stage disease where the oncologist has requested preemptive EGFR mutation testing. Hence there may be a selection bias in this instance towards a clinical suspicion of EGFR mutation, namely younger, female, Asian, non-smokers, with adenocarcinoma histology (8-11). All lung resection specimens were adenocarcinomas, however no trend towards younger or female patients was observed. Unfortunately, further clinical details including ethnicity and smoking status were not available in this study, and given our small study size, a detailed clinicopathological analysis could not be undertaken. However, as lung resection specimens represent the “gold standard” for the quantity of tumor tissue available for mutation testing; this group was selected as the comparator for small biopsy and cytology specimens.
Paired resection and small biopsy or cytology specimens were not available except in one patient, where the result was concordant between cytology and resection specimens. Hence, we have used mutation rate as a surrogate for the adequacy of the specimen for mutation testing. Specifically, the EGFR mutation rate was of primary concern, given its clinical relevance and indication as the main purpose for DNA mutation testing.
Although the EGFR mutation rate was highest in lung resection specimens, there was no statistically significant difference compared to small biopsy and cytology specimens. All specimen types had an EGFR mutation rate within the published range of 10-40% (8-11,15).
In contrast to EGFR mutations, KRAS mutations are more often seen in Caucasians, males, and smokers (10,11). The KRAS mutation rate was significantly lower in cytology specimens compared to other specimen types. This may be related to selection bias, which may also contribute to the slightly higher EGFR mutation rate in cytology specimens compared to small biopsy specimens, though this was not statistically significant. Our study was also limited by its small size.
While previously considered an inferior specimen for mutation testing, to be avoided in favor of small biopsy specimens if possible (16), our data adds to the recent evidence for the suitability of cytology specimens for EGFR mutation testing. Studies have shown that EGFR mutations can be detected in a variety of cytology specimens, including from smears, cell suspensions, and cell blocks, tested on various platforms, as reviewed by Ellison and colleagues (17). This is reflected in the recent molecular testing guidelines released by the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology (12), acknowledging the suitability of cytology specimens, with a preference for cell blocks. EGFR mutation rates have been found to be comparable between cytology and non-cytology specimens (18-23). Some studies have suggested that cytology specimens may indeed be better than histology specimens, with reports of lower test failure rates in cytology specimens in some instances (19,23). Smouse and colleagues (18) found a significantly higher EGFR mutation rate in cytology compared to surgical pathology specimens, though they acknowledged the possible role of selection bias in their study, which was retrospective like ours.
Cytology specimens may have some advantages over small biopsy specimens for mutation testing. Procedures to obtain a cytology specimen may be less invasive with less attendant risks of bleeding and infection, compared with core needle biopsy specimens, or it may be a therapeutic procedure, such as in draining pericardial or pleural fluid. The presence of tumor cells in the specimen can often be verified immediately, through rapid on site evaluation by a cytologist or cytopathologist, increasing the likelihood of an adequate cell block, which is the recommended cytological preparation for mutation testing (12). Production of a cell block by cytology staff results in a defined period of fixation, whereas the small biopsy specimen may, in the real world, have variable duration of fixation, including for prolonged periods, depending on specimen pick up and laboratory work flow, increasing the risk of DNA fragmentation. The cytology specimen itself generally offers a more pure source of tumor, with less admixed stroma or inflammation than commonly seen in small biopsy specimens. Although we found no significant difference in the mean percentage of tumor cells between cytology and small biopsy specimens, the higher mean percentage of tumor cells in cytology specimens with an EGFR mutation compared to those without highlights the importance of careful attention to tumor cell content.
Our study supports the growing evidence for the utility of cytology specimens for mutation testing, and suggests that they are not inferior to small biopsy specimens. In our study, EGFR mutations can be detected at a frequency consistent with published rates in both cytology and small biopsy specimens. With the growing expectation that cytology and small biopsy specimens offer both a diagnosis and a source of tissue for mutation testing, specimen selection and triage is increasingly vital. Each laboratory should be aware of the limitations of their testing method, and consider each case on an individual basis with particular reference to DNA quality and quantity. Communication between the laboratory, pathologists, and physicians is vital both to determine the availability of specimens and the limitations of the assay when only a small specimen is available.
Acknowledgements
Wendy A. Cooper and Sandra A. O’Toole have received funding from National Foundation for Medical Research and Innovation. Sandra A. O’Toole also gratefully acknowledges support from Sydney Catalyst, Cancer Institute NSW, and philanthropic donations from Mr. David Paradice, The TAG Family Foundation, and ICAP.
Disclosure: The authors declare no conflict of interest.
References
- International Agency for Research on Cancer. GLOBOCAN 2012: Estimated Cancer Incidence, Mortality and Prevalence Worldwide in 2012. Available online: http://globocan.iarc.fr/Pages/fact_sheets_cancer.aspx
- Molina JR, Yang P, Cassivi SD, et al. Non-small cell lung cancer: epidemiology, risk factors, treatment, and survivorship. Mayo Clin Proc 2008;83:584-94. [PubMed]
- Lynch TJ, Bell DW, Sordella R, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med 2004;350:2129-39. [PubMed]
- Paez JG, Jänne PA, Lee JC, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004;304:1497-500. [PubMed]
- Pao W, Miller V, Zakowski M, et al. EGF receptor gene mutations are common in lung cancers from "never smokers" and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci U S A 2004;101:13306-11. [PubMed]
- Mok TS, Wu YL, Thongprasert S, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 2009;361:947-57. [PubMed]
- Rosell R, Carcereny E, Gervais R, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 2012;13:239-46. [PubMed]
- Sharma SV, Bell DW, Settleman J, et al. Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 2007;7:169-81. [PubMed]
- Cheng L, Alexander RE, Maclennan GT, et al. Molecular pathology of lung cancer: key to personalized medicine. Mod Pathol 2012;25:347-69. [PubMed]
- Cooper WA, Lam DC, O'Toole SA, et al. Molecular biology of lung cancer. J Thorac Dis 2013;5:S479-90. [PubMed]
- Lovly C, Horn L, Pao W. Molecular Profiling of Lung Cancer. My Cancer Genome 2014. Available online: http://www.mycancergenome.org/content/disease/lung-cancer
- Lindeman NI, Cagle PT, Beasley MB, et al. Molecular testing guideline for selection of lung cancer patients for EGFR and ALK tyrosine kinase inhibitors: guideline from the College of American Pathologists, International Association for the Study of Lung Cancer, and Association for Molecular Pathology. J Thorac Oncol 2013;8:823-59. [PubMed]
- Thomas RK, Baker AC, Debiasi RM, et al. High-throughput oncogene mutation profiling in human cancer. Nat Genet 2007;39:347-51. [PubMed]
- Jurinke C, Oeth P, van den Boom D. MALDI-TOF mass spectrometry: a versatile tool for high-performance DNA analysis. Mol Biotechnol 2004;26:147-64. [PubMed]
- Yip PY, Yu B, Cooper WA, et al. Patterns of DNA mutations and ALK rearrangement in resected node negative lung adenocarcinoma. J Thorac Oncol 2013;8:408-14. [PubMed]
- Pirker R, Herth FJ, Kerr KM, et al. Consensus for EGFR mutation testing in non-small cell lung cancer: results from a European workshop. J Thorac Oncol 2010;5:1706-13. [PubMed]
- Ellison G, Zhu G, Moulis A, et al. EGFR mutation testing in lung cancer: a review of available methods and their use for analysis of tumour tissue and cytology samples. J Clin Pathol 2013;66:79-89. [PubMed]
- Smouse JH, Cibas ES, Jänne PA, et al. EGFR mutations are detected comparably in cytologic and surgical pathology specimens of nonsmall cell lung cancer. Cancer 2009;117:67-72. [PubMed]
- Pang B, Dettmer M, Ong CW, et al. The positive impact of cytological specimens for EGFR mutation testing in non-small cell lung cancer: a single South East Asian laboratory's analysis of 670 cases. Cytopathology 2012;23:229-36. [PubMed]
- Aisner DL, Deshpande C, Baloch Z, et al. Evaluation of EGFR mutation status in cytology specimens: an institutional experience. Diagn Cytopathol 2013;41:316-23. [PubMed]
- Malapelle U, Bellevicine C, De Luca C, et al. EGFR mutations detected on cytology samples by a centralized laboratory reliably predict response to gefitinib in non-small cell lung carcinoma patients. Cancer Cytopathol 2013;121:552-60. [PubMed]
- Leslie C, Giardina T, Carrello A, et al. Detection of EGFR mutational profile by direct dideoxy sequencing in cytology and non-cytology biopsy samples. Pathology 2014;46:283-8. [PubMed]
- Shiau CJ, Babwah JP, da Cunha Santos G, et al. Sample features associated with success rates in population-based EGFR mutation testing. J Thorac Oncol 2014;9:947-56. [PubMed]


