
The contribution of hereditary cancer-related germline mutations to lung cancer susceptibility
Introduction
The germline mutations in multiple genes confer significant risks to several cancers, including breast, ovarian, colorectal cancer and melanoma. In contrast, the genetic predisposition of lung cancer has not yet been elucidated. Although most lung cancers develop sporadically and cigarette smoking is considered to be the predominant risk factor (1), many lung cancer patients present a family clustered pattern. It was reported that a family history confer a substantial risk to lung cancer, especially for those with two or more affected individuals in a family (2).
Since the incidence of definite pathogenic germline mutations are very low, most studies on germline mutations in lung cancer were case report studies, and only a couple of population-based studies so far reporting the prevalence of germline mutations in lung cancer (3-5). Germline EGFR mutations are by far the most frequently reported genetic variations in lung cancer (6), among which EGFR T790M was the most reported germline mutation. It was reported that the prevalence of EGFR T790M germline mutations in East Asian was much lower than that in the Western population (7-9). Therefore, the germline mutation spectrum in lung cancer in different ethnics may be distinct. Other EGFR germline mutations, including V843I, R776G/H, P848L, K757R, D1014N, I646S, G724S, V786M, L792F, R831H, and L844V were also reported with very low incidence (7-9). Apart from EGFR, germline mutations of other genes, including HER2, RET, BRCA1, BRCA2 (9), PARK2 (10), YAP1 (11), CHEK2 (12), TERT (13), TP53, CDKN2A, MET, NBN (14), were also reported and linked with lung cancer risk.
Although some germline mutations, such as those in EGFR and HER2, have been identified in lung cancer in previous observations (3-14), the susceptibility of lung cancer with known hereditary cancer-related germline mutations has not been investigated, and the correlation between germline mutations and somatic mutations has not been studied in detail. The information is sorely lacking among the Chinese population. In this study, we studied the potential susceptibility of lung cancer by categorizing the germline mutations of individual lung cancer patients into three groups based on pathogenicity. Germline and somatic mutation spectrum for each group were obtained by next-generation sequencing (NGS) with a 58-gene panel and a 605-gene panel, respectively. Potential risk factors, such as age, sex, family history, and cancer characteristics, such as cancer type, mutation frequency, tumor mutation burden (TMB) and aberrant pathways, were investigated and compared.
Methods
Ethic approval by participating hospitals
All experiment plans and protocols for the study were submitted to the ethics/licensing committees of the named participating hospitals for review and approval before the start of the clinical study, and were approved by the corresponding committees of hospitals, including the Chinese PLA General Hospital, the Fourth Medical Center of the Chinese PLA General Hospital, the Fifth Medical Center of the Chinese PLA General Hospital and the Eighth Medical Center of the Chinese PLA General Hospital. Confirmation of approval for clinical studies was received from the ethics board of the Chinese PLA General Hospital (approval number: S2018-081-02) before the start of the clinical study. Since the study was designed as a retrospectively study and used retrospective samples collected by the above hospitals, no informed consent was required. Patients with pathogenic or likely pathogenic germline mutations were informed the test results. All experiments, methods, procedures and personnel training were carried out in accordance with relevant guidelines and regulations of participating hospitals and laboratories.
Study design, patients and samples
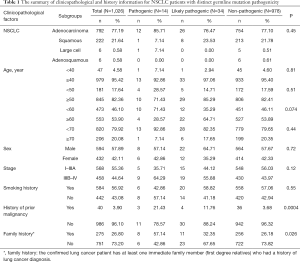
The study was designed and implemented in four Chinese hospitals, and both cancer tissue and blood samples were collected retrospectively. The study was designed to include as many non-small cell lung cancer (NSCLC) patients as possible, as long as the tissue or blood samples were available for next generation sequencing (NGS). As a result, samples collected between June, 2018 and June, 2019 from 1,026 NSCLC patients were obtained based on the availability of samples for NGS test in the participating hospitals, including 792 patients with adenocarcinoma (ADC), 222 patients with squamous cell carcinoma (SCC), 6 patients with large cell carcinoma (LCC) and 6 patients with adenosquamous carcinoma (ASC) (Table 1). Information on clinicopathological status of all patients was collected (Table 1). Family history here is defined as: the confirmed lung cancer patient has at least one immediate family member (first degree relatives) who had a history of lung cancer diagnosis. The immediate family member includes father, mother, brother(s), sister(s), son(s), daughter(s). The collected samples involved tissue samples, including formalin-fix paraffin-embedded (FFPE) samples or frozen samples from surgery or needle biopsy, and blood samples obtained at the time of confirmed lung cancer diagnosis. All technicians were blinded to the clinical information of subjects. The classification of all conditions was based on diagnosis from imaging examinations and subsequent pathological examinations. None of the subjects received chemotherapy, radiotherapy, targeted therapy or immunotherapy before tissue or blood samples were collected. The somatic sequencing data presented in this study were from FFPE samples or frozen tissue samples. Germline sequencing data was obtained from the corresponding genomic DNA of white blood cells.
Full table
Sample preparation, targeted NGS and data processing
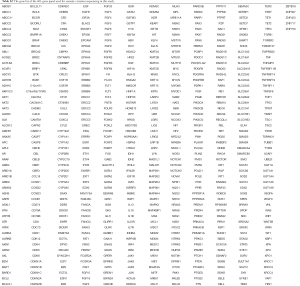
For the FFPE samples, ten 5 µm tumor slices were used for DNA extraction using the QIAamp DNA FFPE Kit (QIAGEN, Valencia, CA, USA) following the manufacturer’s instructions. For blood samples, 2 mL blood were collected in tubes containing EDTA and centrifuged at 1,600 ×g for 10 min at 4 °C within 2 h of collection. The peripheral blood lymphocyte (PBL) debris was stored at −20 °C until further use. DNA from PBLs was extracted using the RelaxGene Blood DNA system (Tiangen Biotech Co., Ltd., Beijing, China) according to the manufacturers’ instructions. Both cancer tissue and white blood cell genomic DNA was quantified with the Qubit 2.0 Fluorometer and the Qubit dsDNA HS assay kit (Thermo Fisher Scientific, Inc., Waltham, MA, USA) according to manufacturer’s instructions. Fragmented genomic DNA underwent end-repairing, A-tailing and ligation with indexed adapters sequentially, followed by size selection using Agencourt AMPure XP beads (Beckman Coulter Inc., Brea, CA, USA), and DNA fragments were used for library construction using the KAPA Library Preparation kit (Kapa Biosystems, Inc., Wilmington, MA, USA) according to the manufacturer’s protocol. Hybridization-based target enrichment was carried out with HaploX germline gene panel (58 known hereditary cancer-related genes, HaploX Biotechnology, gene list is provided in Table S1) for white blood cell genomic DNA or HaploX pan-cancer gene panel (605 cancer-relevant genes, HaploX Biotechnology, gene list is provided in Table S2) for cancer tissue sequencing. Seven to eight polymerase chain reaction (PCR) cycles, depending on the amount of DNA used, were performed by pre-capture ligation-mediated PCR (Pre-LM-PCR) Oligos (Kapa Biosystems, Inc.) in 50 µL reactions. DNA sequencing was then performed on the Illumina Novaseq 6000 system according to the manufacturer’s recommendations at an average depth of 2,200×.

Full table
Full table
Data which meet the following criteria were chosen for subsequent analysis: the ratio of remaining data filtered by fastq in raw data is ≥85%; the proportion of Q30 bases is ≥85%; the ratio of reads on the reference genome is ≥85%; target region coverage ≥98%; average sequencing depth in tissues is ≥2,200×. The called somatic variants need to meet the following criteria: the read depth at a position is ≥20×; the variant allele fraction (VAF) is ≥2% for tissue and PBL genomic DNA; somatic-P value ≤0.01; strand filter ≥1. VAF were calculated for Q30 bases. The copy number variation (CNV) was detected by CNVkit version 0.9.3 (https://github.com/etal/cnvkit). Further analyses of genomic alterations were also performed, including single nucleotide variants (SNVs), CNVs, insertion/deletion (Indels), fusions and structural variation.
Interpretation of pathogenicity of germline mutations and calculation of somatic TMB
Pathogenicity of germline mutations was defined and predicted based on the five-grade classification system according to the American College of Medical Genetics and Genomics (ACMG) Guidelines for the Interpretation of Sequence (15). The VUS, benign and likely benign mutations were defined as the non-pathogenic group (Non-P) in this study. As a result, all germline mutations were categorized into pathogenic (P), likely pathogenic (LP) or non-pathogenic group (Non-P) in this study. TMB was calculated by dividing the total number of tissue non-synonymous SNP and INDEL variations (VAF >2%) by the full length of the exome region of the 605-gene NGS panel (Table S2). Genomic sequence from the DNA of PBLs was used for genomic alignment when calling the somatic mutations.
Statistics and data analysis
Statistical analysis was performed and figures were plotted with GraphPad Prism 5.0 software (GraphPad Software, Inc, La Jolla, CA 92037, USA). Student t-test was performed when two groups were compared, and ANOVA and post hoc tests were performed when three or more groups were compared. Chi-square test and Fisher test were performed when rate or percentage was compared for significance. Figures for mutation spectrum were made with the R software (https://www.r-project.org/). Data for pathway enrichment analysis was analyzed using the method described by DAVID Bioinformatics Resources 6.8 (https://david.ncifcrf.gov/) and visualized by corresponding packages of the R software. The odds ratio was calculated based on the frequency of a certain germline mutation from the Genome Aggregation Database (gnomAD) in general population or East Asian population and the corresponding frequency of mutation obtained from this study. The odds ratio and 95% confidence interval (CI) for each germline mutation was calculated using the calculation module from the SPSS 17.0 software (IBM China Company Limited, Beijing 100101, China). P<0.05 is statistically significant.
Results
Characteristics of pathogenic and likely pathogenic germline mutations in Chinese lung cancer patients and their impact on lung cancer risk
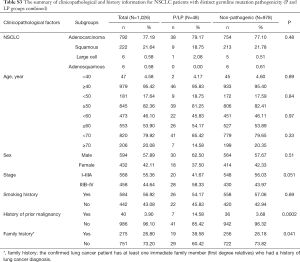
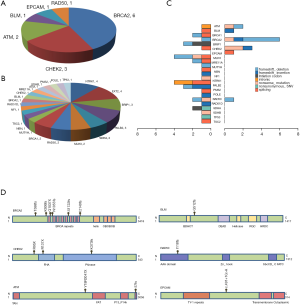
Fourteen patients were found to carry 13 pathogenic (P) germline mutations, and 34 patients carried 36 likely pathogenic (LP) germline mutations, and the remaining 978 patients all carried non-pathogenic (Non-P) mutations (Table 1, Figure 1A,B). No significant difference among the three groups were found with pathological subtypes (P=0.45), age (P values was shown for various age groups in Table 1), stage (P=0.12), sex (P=0.72) or smoking history (P=0.55) (Table 1). This was also true when P and LP groups were combined (Table S3). Interestingly, the ratio of lung cancer patients with at least one immediate family member (first degree relatives) with lung cancer history was significantly higher in the P group than the Non-P group (P=0.009), indicating that pathogenic cancer-predisposing variants predisposed to lung cancer and resulted in familial clustering. Furthermore, the ratio of lung cancer patients with history of other cancers (history of prior malignancy) was higher in P (P=0.0007) or LP (P=0.017) group than the Non-P group (Table 1), suggesting that the presence of pathogenic germline mutations also increased the incidence of other cancers. This was also true when P and LP groups were combined and compared with the Non-P group (Table S3), in which significant differences were also found regarding family history (P=0.041) and history of prior malignancy (P=0.0002).
Full table
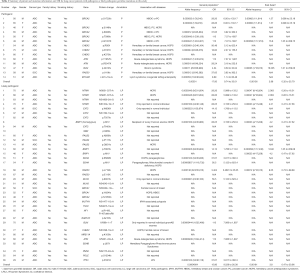
Detailed study identified 6 out of 14 patients in the P group carried BRCA2 pathogenic mutations (6/14), followed by CHEK2 (3/14) and ATM (2/14) (Table 2, Figure 1A). In the LP group, 4 out of 34 patients carried NTRK1 mutations (4/34), 4 carried EXT2 mutations (4/34), followed by BRIP1(3/34) and PALB2 (3/34) (Table 2, Figure 1B). The functions of genes with pathogenic and likely pathogenic mutations mainly involved DNA repair (BRCA1 and BRCA2, BLM, RAD50, BRIP1, MLH3), cell cycle regulation (such as CHEK2, ATM, NTRK1 and EPCAM) and tumor suppressor (such as PALB2 and BRCA1). Most of these fragmental mutations were located within or close to known important protein functional domains (Figure 1C,D) and may have great impacts on protein function.
Full table
In order to study the risk of lung cancer in individuals carrying pathogenic or likely pathogenic germline mutations, we searched the mutation prevalence of all germline mutations in total population and the East Asian population from the Genome Aggregation Database (gnomAD) (Table 2). By comparing the germline mutation frequency found in this study with the variant prevalence in total population and East Asian population, we calculated the overall odds ratio (OR) for the germline mutations in our study. The overall OR value of the P and LP groups was 17.93 (95% CI: 9.74 to 33.01) and 15.86 (95% CI: 5.999 to 133.2), respectively, when compared with the total population, and was 2.88 (95% CI: 0.32 to 25.79) and 3.80 (95% CI: 0.47 to 30.96), respectively, when compared with the East Asian population, suggesting that the pathogenic and likely pathogenic germline mutations were risk factors for lung cancer (Table 2).
Characteristics of somatic mutations of lung cancer patients carrying germline pathogenic or likely pathogenic mutations
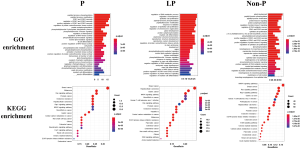
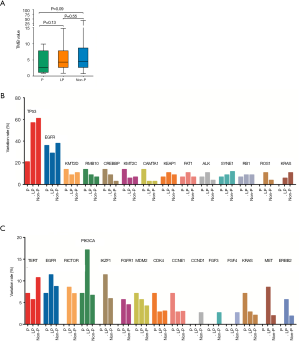
The relationship between germline variations and somatic mutations in lung cancer has not been investigated in detail. We therefore mapped the somatic SNV/INDEL mutation spectrum (Figure S1) and CNV mutation spectrum (Figure S2) categorized by pathogenicity of germline mutations of all lung cancer patients in this study, and investigated the involved genes and somatic mutation characteristics (Figure 2). No statistically significant difference in TMB among the three groups was identified (Figure 2A), however, there was a trend that the TMB in the P group was lower than that of the LP group (P=0.13) and the Non-P group (P=0.09). The average TMB and Inter-Quartile Range (IQR) were 4.07 muts/MB (IQR: 6.74), 5.94 muts/MB (IQR: 5.22) and 6.56 muts/MB (IQR: 6.09) for the P, LP and Non-P group, respectively. The specific driver genes involved attracted our attention. The SNV/INDEL mutation rate (frequency) of TP53 and EGFR was the highest among all genes (Figure 2B). The TP53 mutation rate in the P group was significantly lower than that of the LP (P=0.018) and Non-P groups (P=0.003) (Figure 2B, Figure S1), while no such difference was found with EGFR. We also examined the mutation rate of CNVs in the three groups (Figure 2C). The most common genes with CNVs involved TERT, EGFR, RICTOR and PIK3CA. It appeared that the CNV mutation rate (frequency) of PIK3CA in the LP group was significantly higher than that of the Non-P group (P=0.013) but not the P group (P=0.35) (Figure 2C, Figure S2). Furthermore, the CNV mutation rate of the MET in the LP group was significantly higher than that of the Non-P group (P=0.011). Pathway enrichment analysis on P, LP and Non-P groups was performed, and both GO and KEGG enrichment revealed no significant differences in the functions or biological processes among the P, LP and Non-P groups (Figure S3).


Discussion
Our study provided the first set of evidence on the correlation between the hereditary tumor-related germline mutations and the risk of lung cancer in Chinese population. We found that BRCA2 accounted for the top pathogenic mutations (6/14) in Chinese lung cancer patients, followed by CHEK2 (3/14) and ATM (2/14). Pathogenic mutations were mainly frameshift and nonsense, indicating that germline mutations causing large fragment alterations were the main types in Chinese lung cancer patients. In addition, the functions of BRCA2, CHEK2, ATM, BLM, EPCAM and RAD50 are mainly related to DNA repair and cell cycle regulation, suggesting that the germline mutations of these genes may cause dysregulation of DNA repair and cell cycle and be one genetic risk factor for the development of lung cancer. In the LP group, there were also many splicing mutations in addition to frameshift mutations, indicating that the influence of non-coding splicing sites on protein function cannot be ignored. In this study, the somatic mutations in patients with pathogenic or likely pathogenic germline mutations showed some interesting features. The trend of lower TMB in the pathogenic group indicated the somatic mutations in patients with pathogenic germline variations may be more focused on key driver genes and key pathways, while the somatic mutations in patients without pathogenic germline variations may be more sporadic. Therefore, patients with pathogenic germline mutations may be more likely to develop aberrancies in key driver genes and key pathways, leading to increased risk of lung cancer. It is interesting to find that the affected pathways in patients with or without pathogenic germline mutations were similar, suggesting that the carcinogenesis mechanism of pathogenic group would be consistent with that from the non-pathogenic groups, i.e., the sporadic lung cancer patients, in which cigarette smoke-induced genotoxic damage or other environmental hazards are main causes of malignant transformation (1,2). This indicates that the influence of pathogenic germline mutations mimics the effects of the smoke and environmental factors. One possible explanation for this phenomenon is that the affected germline mutations happen to be those mainly relating to DNA damage and repair. Another possibility is that the presence of pathogenic germline mutations possibly increased the susceptibility to these risk factors and individuals are more likely to develop mutations relating to these factors.
Germline mutations that have been reported in previous studies have focused primarily on EGFR mutations (9,14), mainly because the use of TKI is closely related to EGFR mutations. However, EGFR mutations are not conventional germline mutations related to hereditary cancers, and population studies have reported that EGFR germline mutations were not common in lung cancer [prevalence of 0.13% (12/9,091)] (9), although EGFR germline mutations at multiple sites have been reported (14). Its incidence is even lower in general population with no lung cancer. Therefore, the significance of large-scale screening for EGFR germline mutations in general population is not clear due to its low incidence. However, lung cancer patients and their relatives may benefit from the screening of EGFR germline mutations. In contrast, the BRCA2 germline mutations in this study exhibited a higher overall incidence of 0.68% (7/1,026) than EGFR germline mutations, and therefore may be of more significance in clinical guidance and risk assessment for patients and their families. In addition to EGFR, previous studies have also found that germline susceptibility loci of multiple genes in lung cancer patients were associated with lung cancer risk, including ATM, BRCA2, CHEK2, EGFR, PARK2, TERT, TP53 and YAP1 (5), BRCA1, BRCA2, ERCC4, EXT1, HNF1A, PTCH1, SMARCB1, TP53 (16), BRCA2 p.Lys3326X, CHEK2 p.Ile157Thr, TP63, rs13314271 (12), ARHGEF5, ANKRD20A2, ZNF595, ZNF812, MYO18B (17), and BRCA2 K3326X, LTB p.Leu87Phe, P3H2 p.Gln185His, DAAM2 p.Asp762Gly (18). Among these studies, Parry and colleagues (5) performed a population-based study with TCGA database and found that the ATM gene accounted for 50% of lung cancer germline mutations, followed by TP53, BRCA2, EGFR, and PARK2. This was quite different from the prevalence of germline mutations found in this study, which may be due to the selection of different populations and different target genes. In another recent population-based study, BRCA2 germline mutations ranked the highest in all germline mutations tested, with a detection rate of 0.38% (17/4,459) (3), which was similar to the finding of this study. It should be noted that the above two population-based studies included only 8 or 16 germline genes (3,5). In contrast, our study containing 58 germline genes is therefore more comprehensive and representative than the above studies in reflecting the profile of germline mutations in lung cancer patients.
We found that the somatic average mutation rate varied with different germline mutations. For example, the mutation rate of TP53 in the P group was significantly lower than that of the other two groups, while no such difference in the mutation rate of EGFR was observed, which indicates differential effects of pathogenic germline mutations on somatic driver genes. Interestingly, the CNV mutation rate of PIK3CA and MET of the LP group were significantly higher than that of the Non-P group, suggesting that the somatic amplification of these two genes may be more prominent than other genes when likely-pathogenic germline mutations were present. These observations indicate that the activation of PI3K/AKT and MET pathways may be characteristic in CNV-related alterations. We therefore speculate that patients with DDR-related germline driver gene mutations (such as BRCA2) may be affected by both germline and somatic driver gene mutations, suggesting a different mechanism and a higher risk compared with those without germline driver gene mutations.
The frequency of mutations queried in the GnomAD database represents the frequency of a certain mutation site in the general population. Since most pathogenic or likely pathogenic germline mutations exhibited very low incidence in the general population, the frequency in the database may have certain randomness and may not accurately represent the true frequency in the population. Similarly, the frequency of pathogenic or likely pathogenic germline mutations found in this study was also affected by randomness, and the OR value for a single mutation site may not accurately represent the true frequency in lung cancer population. However, when we pooled all the germline mutations together, the overall mutation frequency was statistically significant, and the overall OR of the P or LP group was comparable with that from the gnomAD database. In this study, the OR of the P group and the LP group suggested that the germline mutations were risk factors for lung cancer. This was also observed in previous studies on lung cancer germline mutations. For example, Parry et al. reported that the overall OR was 66 from 14 germline mutations including ATM and TP53 (5), and Wang et al. reported that the OR for BRCA2 L3326X was 2.47 (12). It is not easy to define the OR value of a certain locus of a certain gene, as the sample size for lung cancer patients and general population need to be large enough for the value to be accurately calculated. Therefore, the report from Parry et al. and our study estimated the overall OR of pooled germline mutations to assess the risk of lung cancer in population (5). In any case, our study and previous studies have demonstrated that pathogenic germline mutations are a risk factor for lung cancer.
It is not uncommon to see lung cancer patients with a familial history. We identified 26.74% of lung cancer patients in this study who had at least one immediate family member with lung cancer. However, unlike other hereditary tumors, most of these lung cancer patients did not had clear pathogenic germline mutations, and the germline mutations or susceptibility loci of the families reported in the previous cases varied greatly, and no clear genetic abnormalities or aggregation has been identified (17,19,20). Therefore, it can be speculated that the occurrence of familial lung cancer may be due to a combination of multiple genetic factors and environmental factors. Elucidation of these factors may require comprehensive family study including typical familial lung cancer patients and their relatives to collect enough data for correlation analysis. In contrast, familial risk is relatively clear for lung cancer patients with clear pathogenic or likely pathogenic germline mutations, therefore, screening for germline mutations in lung cancer patients can help their relatives to understand the risk of the disease and prevent it in advance. Meanwhile, due to the high proportion of BRCA2 pathogenic germline mutations in Chinese population, PARP inhibitors may be applied for this specific population in addition to traditional chemoradiotherapy, targeted therapy or immunotherapy, and relevant clinical trials have also shown positive results (21). Future studies on germline mutations in lung cancer patients should focus on the identification of genetic factors of familial lung cancer and the elucidation of pathogenicity of germline mutations, which will help more patients and their relatives with the prevention and treatment of lung cancer.
Acknowledgments
Funding: This study was supported by the Special Funds for Strategic Emerging Industry Development of Shenzhen (grant number 20170922151538732), and the Science and Technology Project of Shenzhen (grant number JSGG20180703164202084). All funders did not participate in the study design, study implementation, data collection, data analysis, data interpretation and manuscript writing of the study.
Footnote
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tlcr-19-403). Mengyuan L, XL, PS, TH, YZ, Ming L, LL, Yaru C, YZ, GL, JY and SC report non-financial support from HaploX Biotechnology outside the submitted work. LS reports grants from The Special Funds for Strategic Emerging Industry Development of Shenzhen and The Science and Technology Project of Shenzhen, non-financial support and other from HaploX Biotechnology Co., Ltd. outside the submitted work. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the ethics board of the Chinese PLA General Hospital (approval number: S2018-081-02) and individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Alberg AJ, Brock MV, Ford JG, et al. Epidemiology of lung cancer: Diagnosis and management of lung cancer, 3rd ed: American College of Chest Physicians evidence-based clinical practice guidelines. Chest 2013;143:e1S-e29S.
- Lissowska J, Foretova L, Dabek J, et al. Family history and lung cancer risk: international multicentre case-control study in Eastern and Central Europe and meta-analyses. Cancer Causes Control 2010;21:1091-104. [Crossref] [PubMed]
- Slavin TP, Banks KC, Chudova D, et al. Identification of Incidental Germline Mutations in Patients With Advanced Solid Tumors Who Underwent Cell-Free Circulating Tumor DNA Sequencing. J Clin Oncol 2018. [Crossref] [PubMed]
- Hu Y, Alden RS, Odegaard JI, et al. Discrimination of Germline EGFR T790M Mutations in Plasma Cell-Free DNA Allows Study of Prevalence Across 31,414 Cancer Patients. Clin Cancer Res 2017;23:7351-9. [Crossref] [PubMed]
- Parry EM, Gable DL, Stanley SE, et al. Germline Mutations in DNA Repair Genes in Lung Adenocarcinoma. J Thorac Oncol 2017;12:1673-8. [Crossref] [PubMed]
- Yamamoto H, Yatabe Y, Toyooka S. Inherited lung cancer syndromes targeting never smokers. Transl Lung Cancer Res 2018;7:498-504. [Crossref] [PubMed]
- Ikeda K, Nomori H, Mori T, et al. Novel germline mutation: EGFR V843I in patient with multiple lung adenocarcinomas and family members with lung cancer. Ann Thorac Surg 2008;85:1430-2. [Crossref] [PubMed]
- Prim N, Legrain M, Guerin E, et al. Germline exon 21 EGFR mutations, V843I and P848L, in non-small cell lung cancer patients. Eur Respir Rev 2014;23:390-2. [Crossref] [PubMed]
- Lu S, Yu Y, Li Z, et al. EGFR and ERBB2 Germline Mutations in Chinese Lung Cancer Patients and Their Roles in Genetic Susceptibility to Cancer. J Thorac Oncol 2019;14:732-6. [Crossref] [PubMed]
- Xiong D, Wang Y, Kupert E, et al. A recurrent mutation in PARK2 is associated with familial lung cancer. Am J Hum Genet 2015;96:301-8. [Crossref] [PubMed]
- Chen HY, Yu SL, Ho BC, et al. R331W Missense Mutation of Oncogene YAP1 Is a Germline Risk Allele for Lung Adenocarcinoma With Medical Actionability. J Clin Oncol 2015;33:2303-10. [Crossref] [PubMed]
- Wang Y, McKay JD, Rafnar T, et al. Rare variants of large effect in BRCA2 and CHEK2 affect risk of lung cancer. Nat Genet 2014;46:736-41. [Crossref] [PubMed]
- Landi MT, Chatterjee N, Yu K, et al. A genome-wide association study of lung cancer identifies a region of chromosome 5p15 associated with risk for adenocarcinoma. Am J Hum Genet 2009;85:679-91. [Crossref] [PubMed]
- Shukuya T, Patel S, Shane-Carson K, et al. Lung Cancer Patients with Germline Mutations Detected by Next-Generation Sequencing and/or Liquid Biopsy. J Thorac Oncol 2018;13:e17-e19. [Crossref] [PubMed]
- Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405-24. [Crossref] [PubMed]
- Donner I, Katainen R, Sipilä LJ, et al. Germline mutations in young non-smoking women with lung adenocarcinoma. Lung Cancer 2018;122:76-82. [Crossref] [PubMed]
- Kanwal M, Ding XJ, Ma ZH, et al. Characterization of germline mutations in familial lung cancer from the Chinese population. Gene 2018;641:94-104. [Crossref] [PubMed]
- Liu Y, Lusk CM, Cho MH, et al. Rare Variants in Known Susceptibility Loci and Their Contribution to Risk of Lung Cancer. J Thorac Oncol 2018;13:1483-95. [Crossref] [PubMed]
- Hsu KH, Tseng JS, Wang CL, et al. Higher frequency but random distribution of EGFR mutation subtypes in familial lung cancer patients. Oncotarget 2016;7:53299-308. [Crossref] [PubMed]
- Shukuya T, Takahashi K. Germline mutations in lung cancer. Respir Investig 2019;57:201-6. [Crossref] [PubMed]
- Gadgeel SM. Targeted Therapy and Immune Therapy for Small Cell Lung Cancer. Curr Treat Options Oncol 2018;19:53. [Crossref] [PubMed]


