Circulating DNA in diagnosis and monitoring EGFR gene mutations in advanced non-small cell lung cancer
Serum biomarkers for non-small cell lung cancer (NSCLC)
NSCLC is still the main cause of cancer related death in males and females across Western countries. It is commonly known that about 50% of NSCLC is diagnosed in advanced stage and for the majority of these patients, even if encouraging data regarding immunotherapy have been published, to date chemotherapy still represents the mainstay of treatment and prognosis remains poor (1,2). However, approximately 15-20% of advanced NSCLC presents a targetable driver mutation, a condition that dramatically changes therapeutic perspectives and patient outcome (3-6).
Mutations in the gene encoding for the epidermal growth factor receptor (EGFR) represent the first driver mutations identified in NSCLC. The presence of the mutation implicates a receptor constitutively activated that continuously gives the cell input favoring proliferation (7,8). In 90% of cases EGFR activating mutations are represented by exon 19 deletions and exon 21 L858R point mutations (9). It has been established that EGFR activating gene alterations are more common in patients with specific clinico-pathological characteristics, such as female, never smoker, Asiatic origin and adenocarcinoma histological subtype. EGFR mutations represent the most important factor for prediction of response to EGFR tyrosine kinase inhibitors (TKIs). In fact, they are associate with significant increase in response rate (approximately 70%) and improvement in progression free and overall survival (OS) (4,5,10). To date, TKI registered as first line therapy for patients with EGFR mutated NSCLC are gefitinib, erlotinib and afatinib and their toxicity profile is certainly more tolerable than standard chemotherapy. However, for drug prescription purpose, the presence of EGFR mutation needs to be demonstrated and therefore neoplastic tissue sample is always required.
Unfortunately, biopsies in lung cancer represent a criticism. Bronchoscopy and trans-thoracic biopsies are not well accepted by patients and the event that tumoral material is not sufficient or adequate for molecular analyses is not so infrequent (11). Bone biopsies are a critical issue because decalcification procedures interfere with molecular testing and results (12). Moreover, a single biopsy cannot reflect the clonal heterogeneity of the tumor, which could be present in a single tumor lesion (intratumoral heterogeneity) or between different sites of the tumor (intermetastatic heterogeneity) (13-15). Finally, bioptic procedures are not free from related risks (16). Recent advances in therapeutic management of patient with EGFR mutated NSCLC demonstrated the importance of identifying, after the progression to TKI, the molecular mechanisms of acquired resistance in order to continue, as long as possible, a tailored therapy based on the developed resistance alteration (17,18). This approach entails the repetition of a biopsy theoretically every time a patient experiences a progression of disease with a consequent increased discomfort for the patient who undergoes re-biopsy. Moreover, the re-biopsy after progression is not feasible when disease progression involves a body site that can be reached only with complicated surgical procedures (i.e., brain). All this considerations have given the research the incentive for the identification of more accessible and tolerated methodologies for molecular alteration identification.
Several attempts were done in order to identify reliable serum biomarkers for cancer. In the past, serum proteins, such as for example carcinoma carcinoembryonic antigen (CEA), have been commonly used for diagnosis of different cancer but due to low specificity and sensibility their routinely use is not recommended (19). Subsequently, the identification of circulating tumor cells (CTCs) in serum of patient with cancer seemed to represent the solution for cancer serum diagnosis and monitoring. However, several problems emerged regarding the best method for their isolation as different available devices, basing the selection on cells dimension or antigen expression, presented a moderate risk of false negatives (20). Recently, the attention moved to the possibility of isolation and analysis of cell-free tumor DNA (cftDNA) that, to date, represents the best candidate for identification and monitoring of molecular tumor-related alterations in blood of patients with cancer (21).
Fragments of circulating DNA were isolated in plasma many years ago (22). In particular, patients with cancers present higher levels of circulating DNA comparing to healthy volunteers because of the presence of tumoral counterpart, which express the same molecular abnormalities expressed by DNA of primitive mass (13). The elevate cellular turnover and consequent cellular necrosis and apoptosis cause a massive release of tumoral DNA into the bloodstream were it can be isolated and analyzed. Therefore, tumor size, localization and vascularity may influence cftDNA plasmatic levels. It is also possible that part of cftDNA comes from CTCs lysis (13). The analysis of cftDNA, defined as liquid biopsy, could be repeated every time needed and without any discomfort for patients. Moreover, the mutational analysis of cftDNA demonstrated a significantly better sensitivity if compared with CTCs one, establishing cftDNA as the best circulating source for molecular analysis (23). Information derived from liquid biopsy could be used in future for early cancer diagnosis, assessment of genetic determinants for targeted therapies, monitoring of tumor dynamics and early evaluation of tumor response, identification of resistance mechanisms (13).
In the last years, techniques for cftDNA analysis have been largely employed for identification of activating and resistance mutations in NSCLC EGFR mutated patients and the aim of this review is discuss principal findings.
Circulating free tumor DNA and technologies for its detection
cftDNA could be a relevant biomarker to molecular diagnosis and monitor treatment resistance, because of its sensitivity and specificity, but it really needs reproducible and standardized methods, both for the extraction and for its analyses.
Most of the published papers used conventional methods for the cftDNA extraction with commercially available kits for routine use, based on selective binding to a silica-based membrane for improved recovery of fragmented nucleic acids (i.e., Qiagen, Norgen). While the amount and the quality of cftDNA can deeply vary, high-analytical sensitivity and specificity techniques are required for its detection; moreover, a critical issue is to make a distinction and a choice between the importance and the clinical role of cftDNA quantification and mutation analysis. Because of it, many published studies applied a combined quantitative and qualitative analysis of cftDNA starting from surgery and during follow-up, founding that during follow-up, cftDNA levels decrease progressively, but rapidly increased when a relapse occurred, whereas specific mutations were detected only in relapsed patients (24). Dawson and colleagues analyzed the cftDNA of 30 metastatic breast cancer patients to monitor response to treatment. cftDNA was detected in 29/30 patients, showing that cftDNA levels have a dynamic range and the correlation with variations in tumor burden were better than did CA 15.3 serum biomarker or CTCs (25).
Regarding the mutation analysis of cftDNA, a large number of technologies is now available to analyze mutations in cftDNA, including automatic sequencing, real-time polymerase chain reaction (PCR) platforms, mass spectrometry (MS) genotyping, amplification protocols with magnetic beads in oil emulsions [beads, emulsion, amplification and magnetics (BEAMing)] and next-generation sequencing (NGS), digital PCR platforms (26-30). The sensitivity range of the available techniques varies from 15% to 0.01%, but one of the major gaps in this field is the lack of standardization of techniques, in order to understand how those techniques are cost-effective and reliable to fit clinical needs.
Among techniques most of them are able to detect mutant allele frequencies with a sensitivity of at least 2%, other, like cold-PCR, can reach somatic mutations at very low frequencies of 0.1-0.5%, and many genotyping approaches can be combined with it to analyze known mutations [i.e., MS-based matrix-assisted laser desorption ionization time-of-flight (MALDI-TOF) genotyping technologies] (31).
Real-time PCR
One of the widely used methods to detect known mutations is the real-time PCR. The real-time PCR works with either TaqMan probes or TaqMan Detection Mutation Assay. TaqMan probes have a sensitivity detection limit of approximately 10% (32), otherwise, TaqMan Detection Mutation Assay is a competitive allele-specific TaqMan PCR technology, with high sensitivity and specificity because the mutant allele detection is based on an allele-specific primer, while an MGB blocker oligonucleotide suppresses the wild-type background and high sensitivity. Thanks to this mechanism of action, the TaqMan Detection Mutation Assay is able to detect as low as 0.1% mutant molecules in a background of wild type genomic DNA (Cancer Biomarker Research using castPCR™ Technology, AACR 2012). Real-time PCR can also work with Scorpion primers, a kind of bi-functional molecule in which a primer is covalently linked to the probe, with a fluorophore and a quencher. In the absence of the mutation, the quencher close to the fluorophore absorbs its fluorescence. During the Scorpion PCR reaction, the presence of a mutation separates the fluorophore and the quencher increasing the emitted fluorescence (http://www.premierbiosoft.com/tech_notes/Scorpion.html).
Most of the published studies adopted this technology for the analysis of cftDNA in lung cancer. In particular, results coming from analyses focused on the detection of the EGFR mutations in cftDNA of patients with EGFR mutated tumors showed a wide variability: the concordance ranges from 43% to 100% (23,33). Unfortunately, in some cases, mutations can be missed using Real Time technology, and therefore the results are inferior compared to more sensitive approaches.
Digital PCR
The digital PCR approach is based on the same principle of the real-time PCR, but while the real-time PCR works as a unique solution, the digital PCR is able to divide the amplification mix in several thousand of replicates. This partition permits the amplification and the analysis considering single spots, which means that the system is able to decrease the ration of cftDNA/germinal DNA, increasing sensitivity. Digital PCR can works on various principles, for example, silicon chips (Quant studio 3D, Life technologies, Carlsbad, CA, USA) or micro droplets (Bio-Rad Qx100, BioRad, Hercules, CA, USA). This kind of technology can theoretically increase the sensitivity to 1:100,000 molecules of cftDNA in a germinal DNA background (34). Disadvantage of this technology is the not standardized threshold to establish the presence and the amount of mutations.
Beads, emulsion, amplification and magnetics (BEAMing)
Many other approaches, like BEAMing technology, are able to detect a very small amount of mutant DNA sequences in a larger pool of fragments containing wild-type DNA, in order of a single mutant allele in a background of 10,000 wild-type alleles, and it is able to enabling copy-number quantification (35). BEAMing is a sensitive method to detect known genetic mutations, even when at very low copy numbers. The technique is based on a combination of emulsion digital PCR and flow cytometry, with beads, emulsification, amplification and magnetics to achieve the necessary level of sensitivity. DNA sequences are amplified via emulsion PCR covalently bound to magnetic microbeads via streptavidin-biotin interactions; the PCR products generated in each emulsion droplet will remain physically affixed to the microbeads at the end of the reaction, allowing them to be easily separated and purified using a magnet, to determine the presence and number of known mutant variations. The wild-type or mutant DNA can be easily differentiated using flow cytometry. Unfortunately, the BEAMing workflow results complex limiting the feasibility and reproducibility of the technology.
Next-generation sequencing (NGS)
All the mentioned techniques are able to find only known mutations in samples, and this means that a patient need to have a tumor biopsy screened in advance to capture the mutational status, consequently, in terms of costs and standardization of the analysis, it is need to personalized a panel test for each patient. The analysis of cftDNA using NGS technology has recently demonstrated to offer increase detection sensitivity, showing also a good specificity in patients with advanced cancers (27). Published studies demonstrate that deeper sequencing of plasma DNA may allow the problem of clonal heterogeneity and selection (36).
Many NGS technologies are available to date, all of them produce short sequences from single molecules of DNA and it is compared to a reference sequence, allowing the sequencing of large portion of the genome. Selecting only a limited number of sequences of frequently mutated genes, it is easy to reach very deep cover- age of sequencing for candidate mutation loci. This allows the identification of mutated alleles even if highly diluted. Moreover, one of the advantages of the NGS is that whole-genome sequencing of cftDNA can allow the identification of rearrangement and region of copy number aberrations, not detectable with other techniques (27).
Unfortunately, in terms of daily application into the clinic, the use of a NGS technique is still so far, the management of the data requires expert biologists in library preparation, a dedicated bioinformatics support is recommended to solve computational problems that occur during the project and it is an expensive technique.
Genotyping MS
A considerable number of technologies are available for the detection of mutations using MS, but nowadays, the MALDI-TOF MS has become the most used method. The genotyping method is able to distinguishing different alleles by the different masses of primer extension products. The experimental procedure is divided into three steps: amplification, primers extension reaction, transfer of the reaction product into a chip that contains a specific matrix, with two intermediate cleaning reactions, before detection of the extension products. At the end of the analysis, the peak spectrum resulting from MALDI-TOF MS analysis can be analyzed with software that traces back primer masses to assayed alleles. MALDI-TOF MS is relatively more expensive and time consuming than RT-PCR-based methods, but it is more suitable for the simultaneous analysis of multiple mutations. Sequenom is nowadays into clinical routine for the analysis of somatic mutations from FFPE tissue; one of the limitations of this method, common to other similar genotyping techniques, is that it only returns genotypic data. For this reason, analyses with more than one single nucleotide polymorphism (SNP), such as linkage disequilibrium or haplotype diversity, require the most likely haplotypes to be inferred.
cftDNA for identification of EGFR mutations in patients with NSCLC
To validate cftDNA analysis for EGFR mutations detection, results obtained in serum have been compared with the actual gold standard that is analysis on tissue from tumor biopsy. To our knowledge, the first authors that compared results from serum and paired tissue samples were Kimura and colleagues in 2006 (37). Even if paired samples were just 11, authors reported a 72.7% of concordance between serum and tissue. One year later, the same author published another casistic of 42 patients were EGFR mutational status was consistent with tissue one in 92.9% of cases (38). In 2009, Yung et al. detected EGFR 19del and L858R in 17% and 26%, respectively, of 35 pre-therapy plasma samples by using digital PCR; when data were compared with results from tumor samples, overall serum analysis demonstrated very high sensitivity and specificity (92% and 100%, respectively) (26).
Other studies published and conducted on Asiatic populations, revealed high grade of specificity and moderate grade of sensitivity (39,40). Furthermore, authors observed a significant increase in sensitivity when only patients with advanced stage or poorly differentiated adenocarcinoma were evaluated. These data can be explained considering that overall tumor mass and aggressivity can influence levels of cftDNA and therefore the possibility of EGFR mutation detection.
The principal data regarding Caucasian patients were published by Weber and Douillard (41,42). Weber et al. analyzed pairs of diagnostic biopsy and plasma sample of 199 patients obtained prior commencing therapy with EGFR-TKI (41). The overall concordance between plasma and tissue was 179/199 (90%) and six mutations were present only in plasma sample but not in bioptic specimens suggesting a possible role of tumoral heterogeneity. Douillard and colleagues published data regarding patients enrolled in the phase IV study of gefitinib in Caucasian patients with advanced stage IV EGFR mutated NSCLC (42). All patients were centrally screened for EGFR mutation in tissue sample and matched baseline plasma samples were mandatory. Authors matched 652 tumor and plasma samples and concordance resulted 94.3%, sensitivity 65.7% and specificity 99.8%, concluding that, even if tumor remains the preferred source, plasma testing could be appropriate in patients without available tissue. This statement is based on the evidence that patients with EGFR mutated cftDNA presented a response rate similar to patient with EGFR mutated tissue.
Recently, Mok published results of analysis conducted on data from the FASTACT-2 study where patients were randomized to receive platinum-based chemotherapy plus sequential erlotinib or placebo (43). Authors matched 238 plasma and tissue samples and concordance was 88%, sensitivity 75% and specificity 96%. Similar to previous study, patients with EGFR-positive cftDNA treated with erlotinib presented a significantly better outcome than patients treated with placebo [progression-free survival (PFS) 13.1 vs. 6.0 months; P<0.0001], while no difference emerged between EGFR-negative cftDNA patients treated with erlotinib or placebo. These results enforce the role of cftDNA EGFR mutations as predictive factor for response to EGFR-TKI confirming they could represent a reliable surrogate of tissue determination.
Considering the high number of reports present in literature, two meta-analysis investigating the diagnostic value of cftDNA for EGFR mutations identification have been published and both included studies with paired tissue and plasma samples (44,45). Characteristics of the studies included in the two meta-analyses are summarized in Table 1. The first one considered results from 20 published studies of which all were conducted in Asia but one conducted in USA (44). Results showed a pooled sensitivity of 0.674 (95% CI: 0.517-0.800) and a pooled specificity of 0.935 (95% CI: 0.888-0.963). Positive and negative likelihood ratios were 10.307 (95% CI: 6.167-17.227) and 0.348 (95% CI: 0.226-0.537), respectively. The summary receiver operating characteristic (SROC) curve was generated and area under the curve (AUC) resulted 0.93 [0.90-0.95] indicating high diagnostic accuracy. The other meta-analysis considered 27 studies of which a consistent part already included in the previous one, five studies regarding Caucasian populations and five studies published in 2014 including ones by Douillard and Weber. Pooled sensitivity and specificity were 0.620 (95% CI: 0.513-0.716) and 0.959 (95% CI: 0.929-0.977), respectively and AUC was 0.91 (95% CI: 0.89-0.94). As previously reported, accuracy increased in patients with advanced stage disease (AUC 0.96, 95% CI: 0.94-0.97). The authors of both meta-analyses conclude in favor of the high diagnostic accuracy showed by cftDNA underlying the high specificity and non-invasivity that make it a useful tool for screening. However, some limitations have been described including the presence of heterogeneity between studies and the absence of a unique and specified time of blood collection that could have a significant impact as chemotherapy could influence EGFR status (66).
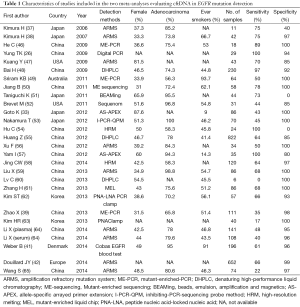
Full table
After publication of these meta-analysis, results of two relevant studies (ASSESS and IGNITE trials) investigating the utility of ctDNA in plasma for the detection of EGFR mutation were presented at European Lung Cancer Conference 2015 (67,68). Both are multicenter diagnostic studies evaluated the utility of ctDNA for EGFR mutation testing in a real-world setting (Europe and Japan in ASSESS and Asia-Pacific and Russia in IGNITE, respectively), having as primary objective the concordance between EGFR mutation status obtained via tissue or cytology and plasma-based testing (Table 2). Both studies have controversial results, probably in relation to heterogeneous methodologies used; in fact, if plasma samples were processed in central designated laboratories, nevertheless EGFR mutation testings on tissue were performed according to local practices and, sometimes, with low sensitive techniques. In ASSESS trial, 1,311 patients were enrolled with data available on both tissue and plasma samples of 1,162. Considering overall results, the concordance obtained was 89.1%, with a sensitivity of 46%, specificity of 97.4%, positive predictive value (PPV) of 77.7% and negative predictive value (NPV) of 90.3%. Considering a subgroup with same methodology used in tissue and plasma, in particular as Therascreen®, results improve (concordance 94.9%, sensitivity of 72.7%, specificity 99.1%, PPV 94.1% and NPV 95%) and are similar those obtained in previous small experiences (42). In IGNITE trial, 3,382 patients were enrolled with data available on both tissue and plasma samples of 2,581. Results obtained (see Table 2) showed findings that need some clarification, in particular in Russian patients; in fact, PPV is low, the percentage of mutations in non-adenocarcinoma is higher than expected (about 10% of cases overall, with higher percentage in plasma than in tissue samples in Russian patients, 7.1% vs. 3.7%, respectively), as well as the percentage of rare mutations (15.5% and 26.7% in Russian adenocarcinoma and non-adenocarcinoma samples, respectively).
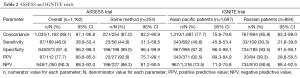
Full table
The role of KRAS mutations in patients with EGFR mutated NSCLC is still controversial. In fact, EGFR and KRAS mutations have always been considered mutually exclusive in lung cancer and KRAS mutations demonstrated a negative predicting effect for response to EGFR-TKI. However, recently studies demonstrating the coexistence of these molecular alterations on tissue samples were published (69). Authors observed that KRAS mutation did not preclude response to EGFR-TKI suggesting that the interaction between the two pathways may be more complex (69). Coexisting EGFR and KRAS mutations have been isolated also in plasma in some studies (62,70,71). Wang et al. reported EGFR/KRAS co-presence in five out of 120 patients who presented PFS and OS significantly inferior to patients harboring only EGFR mutation (70). The presence of both mutations at diagnosis was reported also by Kim et al. in five out of 57 patients. However, in their experience, KRAS serum mutation did not influence prognosis (62). It is worth noting that advances in technologies for DNA molecular analysis could open new scenarios and the role of different mutations may be re-assessed.
Acquired resistance to EGFR-TKI and role of cftDNA
Unfortunately, acquired resistance is an inevitable process during therapy with EGFR-TKI and usually it develops after a median treatment period of 10-12 months (72). Molecular mechanisms underlying acquired resistance have been largely investigated and the occurrence of a second EGFR mutation in exon 20 (T790M) resulted the most frequent resistance-associated molecular alteration with a prevalence ranging from 49% to 63% (72,73). Other less frequent mechanisms of resistance are represented by HER2 amplification (12-13% of cases), MET amplification (5-11%), PIK3CA mutations (about 5%) or BRAF mutations (1%) (73,74). A particular situation is represented by the emergence of a neoplastic clone with clinical and histological features consistent with small cell lung cancer (SCLC) that is reported in 3-14% of cases and implies a more aggressive behavior (72-74).
T790M was reported for the first time in 2005 and its presence increases receptor affinity for ATP that reduces TKI capability to bind EGFR translating in drug inefficacy (75-77). The presence of a clone harboring T790M resistance mutation has been associated with indolent progression and favorable prognosis (78). In fact, Oxnard and colleagues evaluated T790M expression in patients with EGFR-TKI acquired resistance and found out that T790M was significantly more frequent in loco-regional sites of disease than in distant ones and associated with longer post-progression survival. On the contrary, patients without T790M were more likely to progress with new sites of disease in previously uninvolved organs and presented poorer performance status. Similar results have been reported by Oya and colleagues (79); 48% of patients presented T790M in the re-biopsy specimen that was significantly associated with more local than systemic disease progression. Different results were recently reported by Zheng et al. in a Chinese cohort of 117 patients; in fact, even if T790M prevalence (47%) in resistant patients and early onset are confirmed, authors showed that T790M patients presented significantly shorter OS (80).
The importance of the identification of the mechanism involved in acquired resistance is not only theoretical since the efficacy of next generation EGFR-TKI has been demonstrated. Recently, results from trials testing two new molecules AZD9291 and rociletinib have been published and show an impressive efficacy especially in T790M-positive patients, with response rate ranging between 59% and 61% and a median PFS ranging from 9.6 to 13.1 months after progression to first-line TKI (17,18). Similarly to what stated above, the T790M presence need to be demonstrated with re-biopsy after progression and frequently this could represent a limit in lung cancer patients. However, the feasibility of resistance monitoring by plasma DNA sequencing has been proved in several cancers, including EGFR mutated NSCLC [(36), Table 3]. In this study, authors evaluated the variation of mutant allele fractions associated with resistance to oncological treatment in patients with different cancers. Principal findings included the increase of mutations in PIK3CA after therapy with paclitaxel in breast cancer, increase of RB1 mutations after cisplatin in ovarian cancer and increase of T790M in patient with NSCLC EGFR positive treated with gefitinib. T790M was not detectable in plasma at the start of treatment and increased along with NFkB1 and p53 mutations.

Full table
Oxnard et al. reported on a series of nine EGFR mutated patients treated with first-line erlotinib and six of them exhibited T790M in plasma during treatment (81). Sorensen et al. described a group of 23 EGFR mutated patients treated with erlotinib as second-line therapy and the presence of T790M was documented in nine patients as acquired resistance mechanism (82). In particular, authors identified a new response parameter, represented by the plasmatic response, a condition defined by the reduction or disappearance of EGFR activating mutation in plasma during TKI treatment. Reduction in EGFR mutations plasmatic levels can be demonstrated very early, as recently also reported by Marchetti et al., that observed decreased levels starting from the 4th day of therapy with TKI (83). Several authors demonstrated that in patients that developed T790M-mediated acquired resistance, the level of plasmatic EGFR activating mutations started to increase along with the appearance of T790M (81,82,84). Interestingly, in all reports authors demonstrated that T790M was detectable in plasma several days (range: 15-344) before the evidence of disease progression per RECIST criteria. This observation is consistent with the hypothesis of the selection of a resistant neoplastic clone operated by EGFR-TKI, that growths until becomes clinically relevant. However, it should be note that the presence of T790M in association with EGFR sensitizing mutations has been documented in pre-treatment tissue and plasmatic samples, suggesting that the resistance clone could be present since the beginning and reach the blood stream after the clonal expansion (85,91,92). The identification of T790M in patients TKI-naїve could have a significant impact as double-positive patients presented shorter PFS than patients positive only for activating mutations.
Dynamic evolution of EGFR mutation plasmatic levels has been confirmed form others authors. Nakamura et al. reported on a series of 49 patients diagnosed with adenocarcinoma of whom 19 with acquired resistance (86). They found that 53% of resistant patients were positive for T790M and observed that T790M was not detectable in non-responders since T790M appeared in plasma only in responsive patients supporting the theory of a clone selection. Marcq and colleagues described two cases of patients treated with EGFR-TKI (87). In one case activating mutation decreased in plasma and the subsequent increase at progression was associated with T790M appearance; in the other case the patient experienced a complete plasmatic response, with only EGFR activating mutation re-appearing at progression. Wang et al. retrospectively analyzed a series of 135 patients treated with EGFR-TKI and found out that patients with pre-TKI plasma sample positive for T790M had significantly inferior PFS and OS comparing with pre-TKI negative patients (85). Moreover, among patients with pre-TKI positive sample, higher levels were associated with significantly shorter PFS. On the contrary, patients with increased quantity of T790M during TKI therapy presented better PFS and OS than patients with decreasing T790M levels. Interestingly, authors observed high plasmatic levels of MET amplification in patients with decreasing T790M suggesting that TKI pressure could select a MET-amplified tumoral clone responsible of earlier resistance. Similarly to what reported for EGFR activating mutations, also reduction in T790M plasmatic levels can be considered as early parameter of response. In fact, Sequist reported that plasmatic T790M positivity is a predictor of durable response in patients treated with rociletinib, a third generation EGFR-TKI, and that responding patients show decrease of circulating T790M during treatment. However, authors have noted that about 33% of patient with T790M negative plasma responded and that also non-responding patients’ present level reduction during treatment, concluding that probably T790M is not always the dominant resistance driver (89).
Finally, as new third-generation TKI with high affinity for T790M positive receptor have been developed, mechanisms of acquired resistance to new TKI have been studied and identified (88). In a group of 12 re-biopsied patients resistant to rociletinib, Piotrowska and colleagues reported the disappearance of T790M in six patients (of whom two presented transformation to small cell histology) and EGFR amplification in three T790M-positive patients. Regarding plasma analysis, they observed an increased in EGFR activating mutation during TKI therapy that was associated in some patients with T790M increase and in other patients with persistent T790M suppression. Similarly, Thress et al. analyzed plasmatic modifications of patients treated with AZD9291. Together with fluctuations of T790M circulating levels, the appearance of a new mutation C797S was documented as mechanisms of acquired resistance. In vitro studies have documented that this mutation impairs binding of TKI to EGFR thus inducing resistance (90).
Conclusions
Despite tissue biopsy still represents the gold standard for diagnosis, sophisticated technologies have permitted the isolation and identification of lung cancer related mutations in plasma opening new scenarios with a major impact in cancer patients management. Mutational analysis of cftDNA represents one of the most important recent breakthroughs in thoracic oncology. In fact, in certain situations, liquid biopsy could be an essential tool for clinicians because it gives the chance of a targeted therapy also in patients who cannot undergo invasive diagnostic procedures, due to comorbidities or the absence of biopsable tumor lesions. Moreover, liquid biopsy presents the advantages of a non-invasive technique that, without any discomfort, can be repeated every time needed during a patient therapeutic history. In particular, cftDNA analysis assumes a crucial role for patients with EGFR mutated lung cancer, since they represent a group of patients receiving a huge benefit from targeted mutation identification, not only at diagnosis but also at the onset of acquired resistance, but for whom obtaining tissue sample is sometimes not feasible.
Several issues remain outstanding regarding the routine employment of cftDNA. First, many devices for cftDNA detection and analysis have been developed, characterized by a slight different spectrum of sensitivity and specificity. Data in literature are extremely heterogeneous from this point of view as different authors tested the reliability of different devices. Therefore, univocal conclusions cannot still be formulated and two meta-analyses were conducted to clarify the feasibility of plasmatic EGFR mutation detection. Many studies were included, even though conducted with different methods, and globally emerged that plasmatic molecular analysis of EGFR presents a high accuracy suggesting its possible employment when tissue is not available. The evidence that the predictive role of plasmatic EGFR mutation has been confirmed and is consistent with data obtain from tissue enforces the utility of plasmatic analysis for EGFR mutations detection lung cancer. However, diagnostic sensitivity and specificity are influenced also by plasmatic cftDNA levels that depend on cftDNA mechanisms of release and clearance. Moreover, it has been demonstrated that the levels of cftDNA are also determined by several tumor-related factor including tumor mass, stage of disease, vascularization, aggressivity and certainly other are unknown. These issues need to be clarified before cftDNA enter in current clinical practice.
In a minority of patients, the analysis on cftDNA permitted the isolation of KRAS mutation along with the presence of EGFR activating mutation. This is an element of particular interest, as these two alterations have been always considered mutually exclusive and only one report signaled their co-existence in tissue. This finding could be explained considering that plasmatic molecular characterization overcome the limit of tumoral heterogeneity and theoretically permit to identify mutations expressed by clones situated in different body sites. However, it should be considered that new technologies present higher sensitivity than previous ones and therefore could be able to detect molecular alterations expressed by limited number of tumoral cells opening new perspectives on tumor biology.
Finally, the application of cftDNA analysis in the field of acquired resistance to EGFR-TKI is of particular interest. In general, the profile of acquired resistance mechanisms expressed in plasma is consistent to what revealed in tissue samples and T790M, which represent a predicting factor of response to third-generation TKI, emerged as the most frequent resistance mutation. The opportunity of obtaining molecular information avoiding serial re-biopsies permitted to explore the dynamic process leading to resistance. Different authors demonstrated that levels of EGFR activating mutation promptly decreased in plasma after the initiation of EGFR-TKI and that the occurrence of T790M is an early phenomenon that anticipates of several weeks the radiological progression. Again, modifications of T790M levels in response to third-generation EGFR-TKI have been described, even if predictive and prognostic impact is unclear. To date, these findings have not any clinical consequences. However, the efficacy of TKI-therapy modulation basing on fluctuations of plasmatic activating and resistance mutations levels deserved to be valuated prospectively in the future and represent a promising research topic.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Brahmer J, Reckamp KL, Baas P, et al. Nivolumab versus Docetaxel in Advanced Squamous-Cell Non-Small-Cell Lung Cancer. N Engl J Med 2015;373:123-35. [PubMed]
- Reck M, Popat S, Reinmuth N, et al. Metastatic non-small-cell lung cancer (NSCLC): ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 2014;25 Suppl 3:iii27-39. [PubMed]
- Chan BA, Hughes BG. Targeted therapy for non-small cell lung cancer: current standards and the promise of the future. Transl Lung Cancer Res 2015;4:36-54. [PubMed]
- Mok TS, Wu YL, Thongprasert S, et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med 2009;361:947-57. [PubMed]
- Rosell R, Carcereny E, Gervais R, et al. Erlotinib versus standard chemotherapy as first-line treatment for European patients with advanced EGFR mutation-positive non-small-cell lung cancer (EURTAC): a multicentre, open-label, randomised phase 3 trial. Lancet Oncol 2012;13:239-46. [PubMed]
- Solomon BJ, Mok T, Kim DW, et al. First-line crizotinib versus chemotherapy in ALK-positive lung cancer. N Engl J Med 2014;371:2167-77. [PubMed]
- Salomon DS, Brandt R, Ciardiello F, et al. Epidermal growth factor-related peptides and their receptors in human malignancies. Crit Rev Oncol Hematol 1995;19:183-232. [PubMed]
- Zhang X, Gureasko J, Shen K, et al. An allosteric mechanism for activation of the kinase domain of epidermal growth factor receptor. Cell 2006;125:1137-49. [PubMed]
- Shigematsu H, Lin L, Takahashi T, et al. Clinical and biological features associated with epidermal growth factor receptor gene mutations in lung cancers. J Natl Cancer Inst 2005;97:339-46. [PubMed]
- Yang JC, Wu YL, Schuler M, et al. Afatinib versus cisplatin-based chemotherapy for EGFR mutation-positive lung adenocarcinoma (LUX-Lung 3 and LUX-Lung 6): analysis of overall survival data from two randomised, phase 3 trials. Lancet Oncol 2015;16:141-51. [PubMed]
- Vanderlaan PA, Yamaguchi N, Folch E, et al. Success and failure rates of tumor genotyping techniques in routine pathological samples with non-small-cell lung cancer. Lung Cancer 2014;84:39-44. [PubMed]
- Singh VM, Salunga RC, Huang VJ, et al. Analysis of the effect of various decalcification agents on the quantity and quality of nucleic acid (DNA and RNA) recovered from bone biopsies. Ann Diagn Pathol 2013;17:322-6. [PubMed]
- Diaz LA Jr, Bardelli A. Liquid biopsies: genotyping circulating tumor DNA. J Clin Oncol 2014;32:579-86. [PubMed]
- Gerlinger M, Rowan AJ, Horswell S, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med 2012;366:883-92. [PubMed]
- Vogelstein B, Papadopoulos N, Velculescu VE, et al. Cancer genome landscapes. Science 2013;339:1546-58. [PubMed]
- Overman MJ, Modak J, Kopetz S, et al. Use of research biopsies in clinical trials: are risks and benefits adequately discussed? J Clin Oncol 2013;31:17-22. [PubMed]
- Jänne PA, Yang JC, Kim DW, et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N Engl J Med 2015;372:1689-99. [PubMed]
- Sequist LV, Soria JC, Goldman JW, et al. Rociletinib in EGFR-mutated non-small-cell lung cancer. N Engl J Med 2015;372:1700-9. [PubMed]
- Ruibal Morell A. CEA serum levels in non-neoplastic disease. Int J Biol Markers 1992;7:160-6. [PubMed]
- Ilie M, Hofman V, Long E, et al. Current challenges for detection of circulating tumor cells and cell-free circulating nucleic acids, and their characterization in non-small cell lung carcinoma patients. What is the best blood substrate for personalized medicine? Ann Transl Med 2014;2:107. [PubMed]
- Bettegowda C, Sausen M, Leary RJ, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med 2014;6:224ra24.
- Mandel P, Metais P. Les acides nucléiques du plasma sanguin chez l’homme. C R Seances Soc Biol Fil 1948;142:241-3. [PubMed]
- Punnoose EA, Atwal S, Liu W, et al. Evaluation of circulating tumor cells and circulating tumor DNA in non-small cell lung cancer: association with clinical endpoints in a phase II clinical trial of pertuzumab and erlotinib. Clin Cancer Res 2012;18:2391-401. [PubMed]
- Frattini M, Gallino G, Signoroni S, et al. Quantitative and qualitative characterization of plasma DNA identifies primary and recurrent colorectal cancer. Cancer Lett 2008;263:170-81. [PubMed]
- Dawson SJ, Tsui DW, Murtaza M, et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med 2013;368:1199-209. [PubMed]
- Yung TK, Chan KC, Mok TS, et al. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non-small cell lung cancer patients. Clin Cancer Res 2009;15:2076-84. [PubMed]
- Forshew T, Murtaza M, Parkinson C, et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med 2012;4:136ra68.
- Lee YJ, Yoon KA, Han JY, et al. Circulating cell-free DNA in plasma of never smokers with advanced lung adenocarcinoma receiving gefitinib or standard chemotherapy as first-line therapy. Clin Cancer Res 2011;17:5179-87. [PubMed]
- Misale S, Yaeger R, Hobor S, et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 2012;486:532-6. [PubMed]
- Liu P, Liang H, Xue L, et al. Potential clinical significance of plasma-based KRAS mutation analysis using the COLD-PCR/TaqMan(®) -MGB probe genotyping method. Exp Ther Med 2012;4:109-112. [PubMed]
- Li J, Wang L, Mamon H, et al. Replacing PCR with COLD-PCR enriches variant DNA sequences and redefines the sensitivity of genetic testing. Nat Med 2008;14:579-84. [PubMed]
- Wilkening S, Hemminki K, Thirumaran RK, et al. Determination of allele frequency in pooled DNA: comparison of three PCR-based methods. Biotechniques 2005;39:853-8. [PubMed]
- Goto K, Ichinose Y, Ohe Y, et al. Epidermal growth factor receptor mutation status in circulating free DNA in serum: from IPASS, a phase III study of gefitinib or carboplatin/paclitaxel in non-small cell lung cancer. J Thorac Oncol 2012;7:115-21. [PubMed]
- Hindson BJ, Ness KD, Masquelier DA, et al. High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 2011;83:8604-10. [PubMed]
- Diehl F, Li M, Dressman D, et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc Natl Acad Sci U S A 2005;102:16368-73. [PubMed]
- Murtaza M, Dawson SJ, Tsui DW, et al. Non-invasive analysis of acquired resistance to cancer therapy by sequencing of plasma DNA. Nature 2013;497:108-12. [PubMed]
- Kimura H, Kasahara K, Kawaishi M, et al. Detection of epidermal growth factor receptor mutations in serum as a predictor of the response to gefitinib in patients with non-small-cell lung cancer. Clin Cancer Res 2006;12:3915-21. [PubMed]
- Kimura H, Suminoe M, Kasahara K, et al. Evaluation of epidermal growth factor receptor mutation status in serum DNA as a predictor of response to gefitinib (IRESSA). Br J Cancer 2007;97:778-84. [PubMed]
- Zhao X, Han RB, Zhao J, et al. Comparison of epidermal growth factor receptor mutation statuses in tissue and plasma in stage I-IV non-small cell lung cancer patients. Respiration 2013;85:119-25. [PubMed]
- Tseng JS, Yang TY, Tsai CR, et al. Dynamic plasma EGFR mutation status as a predictor of EGFR-TKI efficacy in patients with EGFR-mutant lung adenocarcinoma. J Thorac Oncol 2015;10:603-10. [PubMed]
- Weber B, Meldgaard P, Hager H, et al. Detection of EGFR mutations in plasma and biopsies from non-small cell lung cancer patients by allele-specific PCR assays. BMC Cancer 2014;14:294. [PubMed]
- Douillard JY, Ostoros G, Cobo M, et al. Gefitinib treatment in EGFR mutated caucasian NSCLC: circulating-free tumor DNA as a surrogate for determination of EGFR status. J Thorac Oncol 2014;9:1345-53. [PubMed]
- Mok T, Wu YL, Lee JS, et al. Detection and Dynamic Changes of EGFR Mutations from Circulating Tumor DNA as a Predictor of Survival Outcomes in NSCLC Patients Treated with First-line Intercalated Erlotinib and Chemotherapy. Clin Cancer Res 2015;21:3196-203. [PubMed]
- Luo J, Shen L, Zheng D. Diagnostic value of circulating free DNA for the detection of EGFR mutation status in NSCLC: a systematic review and meta-analysis. Sci Rep 2014;4:6269. [PubMed]
- Qiu M, Wang J, Xu Y, et al. Circulating tumor DNA is effective for the detection of EGFR mutation in non-small cell lung cancer: a meta-analysis. Cancer Epidemiol Biomarkers Prev 2015;24:206-12. [PubMed]
- He C, Liu M, Zhou C, et al. Detection of epidermal growth factor receptor mutations in plasma by mutant-enriched PCR assay for prediction of the response to gefitinib in patients with non-small-cell lung cancer. Int J Cancer 2009;125:2393-9. [PubMed]
- Kuang Y, Rogers A, Yeap BY, et al. Noninvasive detection of EGFR T790M in gefitinib or erlotinib resistant non-small cell lung cancer. Clin Cancer Res 2009;15:2630-6. [PubMed]
- Bai H, Mao L, Wang HS, et al. Epidermal growth factor receptor mutations in plasma DNA samples predict tumor response in Chinese patients with stages IIIB to IV non-small-cell lung cancer. J Clin Oncol 2009;27:2653-9. [PubMed]
- Sriram KB, Tan ME, Savarimuthu SM, et al. Screening for activating EGFR mutations in surgically resected nonsmall cell lung cancer. Eur Respir J 2011;38:903-10. [PubMed]
- Jiang B, Liu F, Yang L, et al. Serum detection of epidermal growth factor receptor gene mutations using mutant-enriched sequencing in Chinese patients with advanced non-small cell lung cancer. J Int Med Res 2011;39:1392-401. [PubMed]
- Taniguchi K, Uchida J, Nishino K, et al. Quantitative detection of EGFR mutations in circulating tumor DNA derived from lung adenocarcinomas. Clin Cancer Res 2011;17:7808-15. [PubMed]
- Brevet M, Johnson ML, Azzoli CG, et al. Detection of EGFR mutations in plasma DNA from lung cancer patients by mass spectrometry genotyping is predictive of tumor EGFR status and response to EGFR inhibitors. Lung Cancer 2011;73:96-102. [PubMed]
- Nakamura T, Sueoka-Aragane N, Iwanaga K, et al. Application of a highly sensitive detection system for epidermal growth factor receptor mutations in plasma DNA. J Thorac Oncol 2012;7:1369-81. [PubMed]
- Hu C, Liu X, Chen Y, et al. Direct serum and tissue assay for EGFR mutation in non-small cell lung cancer by high-resolution melting analysis. Oncol Rep 2012;28:1815-21. [PubMed]
- Huang Z, Wang ZJ, Bai H, et al. The detection of EGFR mutation status in plasma is reproducible and can dynamically predict the efficacy of EGFR-TKI. Thorac Cancer 2012;3:334-40.
- Xu F, Wu J, Xue C, et al. Comparison of different methods for detecting epidermal growth factor receptor mutations in peripheral blood and tumor tissue of non-small cell lung cancer as a predictor of response to gefitinib. Onco Targets Ther 2012;5:439-47. [PubMed]
- Yam I, Lam DC, Chan K, et al. EGFR array: uses in the detection of plasma EGFR mutations in non-small cell lung cancer patients. J Thorac Oncol 2012;7:1131-40. [PubMed]
- Jing CW, Wang Z, Cao HX, et al. High resolution melting analysis for epidermal growth factor receptor mutations in formalin-fixed paraffin-embedded tissue and plasma free DNA from non-small cell lung cancer patients. Asian Pac J Cancer Prev 2014;14:6619-23. [PubMed]
- Liu X, Lu Y, Zhu G, et al. The diagnostic accuracy of pleural effusion and plasma samples versus tumour tissue for detection of EGFR mutation in patients with advanced non-small cell lung cancer: comparison of methodologies. J Clin Pathol 2013;66:1065-9. [PubMed]
- Lv C, Ma Y, Feng Q, et al. A pilot study: sequential gemcitabine/cisplatin and icotinib as induction therapy for stage IIB to IIIA non-small-cell lung adenocarcinoma. World J Surg Oncol 2013;11:96. [PubMed]
- Zhang H, Liu D, Li S, et al. Comparison of EGFR signaling pathway somatic DNA mutations derived from peripheral blood and corresponding tumor tissue of patients with advanced non-small-cell lung cancer using liquidchip technology. J Mol Diagn 2013;15:819-26. [PubMed]
- Kim ST, Sung JS, Jo UH, et al. Can mutations of EGFR and KRAS in serum be predictive and prognostic markers in patients with advanced non-small cell lung cancer (NSCLC)? Med Oncol 2013;30:328. [PubMed]
- Kim HR, Lee SY, Hyun DS, et al. Detection of EGFR mutations in circulating free DNA by PNA-mediated PCR clamping. J Exp Clin Cancer Res 2013;32:50. [PubMed]
- Li X, Ren R, Ren S, et al. Peripheral blood for epidermal growth factor receptor mutation detection in non-small cell lung cancer patients. Transl Oncol 2014;7:341-8. [PubMed]
- Wang S, Han X, Hu X, et al. Clinical significance of pretreatment plasma biomarkers in advanced non-small cell lung cancer patients. Clin Chim Acta 2014;430:63-70. [PubMed]
- Bai H, Wang Z, Chen K, et al. Influence of chemotherapy on EGFR mutation status among patients with non-small-cell lung cancer. J Clin Oncol 2012;30:3077-83. [PubMed]
- Reck M, Hagiwara K, Han B, et al. Investigating the utility of circulating-free tumour-derived DNA (ctDNA) in plasma for the detection of epidermal growth factor receptor (EGFR) mutation status in european and japanese patients (PTS) with advanced non-small-cell lung cancer (ANSCLC): ASSESS study. European Lung Cancer Conference, Geneva, Switzerland, 2015;abstr: 229.
- Han B, Tjulandin S, Hagiwara K, et al. Determining the prevalence of EGFR mutations in Asian and Russian patients (pts) with advanced non-small-cell lung cancer (aNSCLC) of adenocarcinoma (ADC) and non-ADC histology: IGNITE study. European Lung Cancer Conference, Geneva, Switzerland, 2015;abstr: 233.
- Benesova L, Minarik M, Jancarikova D, et al. Multiplicity of EGFR and KRAS mutations in non-small cell lung cancer (NSCLC) patients treated with tyrosine kinase inhibitors. Anticancer Res 2010;30:1667-71. [PubMed]
- Wang S, An T, Wang J, et al. Potential clinical significance of a plasma-based KRAS mutation analysis in patients with advanced non-small cell lung cancer. Clin Cancer Res 2010;16:1324-30. [PubMed]
- Del Re M, Landi L, Tiseo M, et al. Association of KRAS mutations in cell-free circulating tumor DNA with occurrence of resistance to TKIs in NSCLC. J Clin Oncol 2014;32:5 suppl: abstr 11056.
- Sequist LV, Waltman BA, Dias-Santagata D, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med 2011;3:75ra26. [PubMed]
- Yu HA, Arcila ME, Rekhtman N, et al. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin Cancer Res 2013;19:2240-7. [PubMed]
- Sacher AG, Jänne PA, Oxnard GR. Management of acquired resistance to epidermal growth factor receptor kinase inhibitors in patients with advanced non-small cell lung cancer. Cancer 2014;120:2289-98. [PubMed]
- Kobayashi S, Boggon TJ, Dayaram T, et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med 2005;352:786-92. [PubMed]
- Pao W, Miller VA, Politi KA, et al. Acquired resistance of lung adenocarcinomas to gefitinib or erlotinib is associated with a second mutation in the EGFR kinase domain. PLoS Med 2005;2:e73. [PubMed]
- Yun CH, Mengwasser KE, Toms AV, et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc Natl Acad Sci U S A 2008;105:2070-5. [PubMed]
- Oxnard GR, Arcila ME, Sima CS, et al. Acquired resistance to EGFR tyrosine kinase inhibitors in EGFR-mutant lung cancer: distinct natural history of patients with tumors harboring the T790M mutation. Clin Cancer Res 2011;17:1616-22. [PubMed]
- Oya Y, Yoshida T, Tanaka K, et al. Association between clinical outcome of first EGFR-TKIs and T790M mutation in NSCLC patients harboring EGFR mutation with acquired resistance to EGFR-TKIs. J Clin Oncol 2015;33:suppl; abstr e19123.
- Zheng Di, Ye X, Zhang M, et al. Association of plasma EGFR T790M ctDNA status with clinical outcome in advanced NSCLC patients with acquired EGFR-TKI resistance. J Clin Oncol 2015;33:suppl; abstr 8080.
- Oxnard GR, Paweletz CP, Kuang Y, et al. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res 2014;20:1698-705. [PubMed]
- Sorensen BS, Wu L, Wei W, et al. Monitoring of epidermal growth factor receptor tyrosine kinase inhibitor-sensitizing and resistance mutations in the plasma DNA of patients with advanced non-small cell lung cancer during treatment with erlotinib. Cancer 2014;120:3896-901. [PubMed]
- Marchetti A, Palma JF, Felicioni L, et al. Early prediction of response to tyrosine kinase inhibitors by quantification of EGFR mutations in plasma of non-small cell lung cancer patients. J Clin Oncol 2015;33:suppl; abstr 8079.
- Ahn MJ, Lee JY, Lim SH, et al. Dynamic serial monitoring of EGFR mutations in plasma DNA samples in EGFR mutant NSCLC patients treated with EGFR TKI. J Clin Oncol 2015;33:suppl; abstr 8078.
- Wang Z, Chen R, Wang S, et al. Quantification and dynamic monitoring of EGFR T790M in plasma cell-free DNA by digital PCR for prognosis of EGFR-TKI treatment in advanced NSCLC. PLoS One 2014;9:e110780. [PubMed]
- Nakamura T, Sueoka-Aragane N, Iwanaga K, et al. A noninvasive system for monitoring resistance to epidermal growth factor receptor tyrosine kinase inhibitors with plasma DNA. J Thorac Oncol 2011;6:1639-48. [PubMed]
- Marcq M, Vallée A, Bizieux A, et al. Detection of EGFR mutations in the plasma of patients with lung adenocarcinoma for real-time monitoring of therapeutic response to tyrosine kinase inhibitors? J Thorac Oncol 2014;9:e49-50. [PubMed]
- Piotrowska Z, Niederst MJ, Karlovich CA, et al. Heterogeneity Underlies the Emergence of EGFRT790 Wild-Type Clones Following Treatment of T790M-Positive Cancers with a Third-Generation EGFR Inhibitor. Cancer Discov 2015;5:713-22. [PubMed]
- Sequist LV, Goldman JW, Wakelee HA, et al. Efficacy of rociletinib (CO-1686) in plasma-genotyped T790M-positive non-small cell lung cancer (NSCLC) patients (pts). J Clin Oncol 2015;33:suppl; abstr 8001.
- Thress KS, Paweletz CP, Felip E, et al. Acquired EGFR C797S mutation mediates resistance to AZD9291 in non-small cell lung cancer harboring EGFR T790M. Nat Med 2015;21:560-2. [PubMed]
- Rosell R, Molina MA, Costa C, et al. Pretreatment EGFR T790M mutation and BRCA1 mRNA expression in erlotinib-treated advanced non-small-cell lung cancer patients with EGFR mutations. Clin Cancer Res 2011;17:1160-8. [PubMed]
- Watanabe M, Kawaguchi T, Isa S, et al. Ultra-Sensitive Detection of the Pretreatment EGFR T790M Mutation in Non-Small Cell Lung Cancer Patients with an EGFR-Activating Mutation Using Droplet Digital PCR. Clin Cancer Res 2015;21:3552-60. [PubMed]


