
Targeting of arachidonic acid-modulated autophagy to enhance the sensitivity of ROS1+ or ALK+ non-small cell lung cancer to crizotinib therapy
Highlight box
Key findings
• For the first time, we demonstrated that arachidonic acid (AA) can contribute to the induction of crizotinib resistance in non-small cell lung cancer (NSCLC) with ALK- and ROS1-fusion, and targeting AA pathways may represent a new treatment strategy for alleviating tyrosine kinase inhibitor (TKI) resistance.
What is known and what is new?
• NSCLC-patients treated with crizotinib frequently experience drug resistance.
• The AA accumulation induced by PTGS2 downregulation may be a key factor in the development of one of the crizotinib resistance mechanisms in NSCLC.
What is the implication, and what should change now?
• This newly discovered mechanism of ALK- and ROS1-TKI resistance may potentially represent an alternative target for overcoming crizotinib resistance in patients treated with NSCLC.
Introduction
Lung cancer is the leading cause of cancer-related death worldwide, and 85% of lung cancers are non-small cell lung cancer (NSCLC) (1). Among patients with NSCLC, over 70% are diagnosed at a late stage, which limits treatment effects and results in poor prognosis. With the approval of tyrosine kinase inhibitors (TKIs) for NSCLC therapy, the survival rates of patients have been significantly improved. Among them, crizotinib is often used in patients with c-ros oncogene 1 (ROS1)-positive or anaplastic lymphoma kinase (ALK)-rearranged NSCLC (2) and has been shown capable of extending progression-free survival in these patients (3,4). Nevertheless, most patients acquire drug resistance during treatment, significantly limiting the therapeutic effect and posing a serious clinical problem in NSCLC therapy (5). Therefore, further improvements in treatment strategies are required. Several mechanisms of crizotinib resistance in ROS1- and ALK-rearranged NSCLC have been reported, including off-target resistance (bypass signal activation), on-target resistance (gene copy number amplification and secondary mutations,) and phenotypical transformation (5-12). Despite extensive research into resistance mechanisms to crizotinib, drug resistance remains prevalent. Therefore, uncovering new resistance mechanisms by relying on preclinical cell models (13) and identifying potential therapeutic targets remain critical challenges.
Autophagy is a highly conserved process in which cells eliminate unnecessary components, such as protein aggregates, lipid vesicles, or damaged organelles, via lysosomal degradation. This process is activated under conditions of nutrient deprivation or cellular stress to promote cell survival, which contributes to tumor development (14,15), cell dormancy (16), chemoresistance (17,18), immunotherapy resistance (19), and resistance to TKIs (20,21). The autophagic pathway is now regarded as a promising target for overcoming cancer drug resistance (22). A recent study has shown that targeting interleukin-6/signal transducer and activator of transcription 3 (IL-6/STAT3) signaling abrogates EGFR-TKI resistance through inhibiting Beclin-1 dependent autophagy (23). Additionally, Xu et al. reported that CD74-ROS1 L2026M mutant contributes to crizotinib resistance in NSCLC by stimulating autophagy via the MEK/ERK pathway (24). Valencia et al. found that inhibiting tumor-protective autophagy enhances the sensitivity of NSCLC to anti-PD-1 therapy (25). These observations might suggest that autophagy may also be implicated in the development of crizotinib resistance in NSCLC cells. However, the underlying mechanisms remain largely unexplored.
Metabolic reprogramming is recognized as a key event during tumor initiation and progression (26). Emerging evidence indicates that metabolic properties of resistant cancer cells differ significantly from those of their parental cells (27). Targeting metabolic reprogramming might help overcome multiple forms of drug resistance in cancer therapy, including resistance to TKIs, PD-1/PD-L1 immunotherapy and antiangiogenic drugs (28-31). Arachidonic acid (AA), a ω-6 polyunsaturated fatty acid (PUFA), is commonly found in membrane phospholipids and is among the most abundant PUFAs in the human body (32). Upon release from the plasma membrane by phospholipases, AA is converted by prostaglandin endoperoxide synthase 1/2 (PTGS1/2, also known as COX-1/2) into prostanoids like prostaglandins and thromboxane A2, which are involved in inflammation and cell proliferation (33). The PTGS2/COX-2 is the inducible form of the enzyme PTGS (induced by inflammatory stimuli, hormones, growth factors), while PTGS1/COX-1 is the constitutive form (34), and is induced by multiple signaling pathway including STAT3 activation (35). Research suggests that AA can either promote or inhibit tumor cell survival by regulating apoptosis or endoplasmic reticulum (ER) stress (36-38). In recent studies, AA was found to trigger autophagy in Sertoli cells (39) and Caenorhabditis elegans (40). However, the role and mechanism of AA in regulating autophagy and drug resistance in cancer remain unclear. In this study, we aimed to clarify how autophagy is implicated in crizotinib resistance in NSCLC cells and to identify effective related strategies for overcoming crizotinib resistance. Indeed, we found that crizotinib induces apoptosis and growth arrest in ROS1+ or ALK+ NSCLC by inhibiting autophagy flux, which leads to the accumulation of autophagosomes. Conversely, in crizotinib-resistant NSCLC cells, there is a reduction in phosphorylated STAT3 (p-STAT3)/PTGS2 expression, resulting in increased levels of AA. Elevated AA promotes autophagy flux and decreases autophagosome accumulation, contributing to crizotinib resistance under drug stress. Moreover, chloroquine (CQ, an inhibitor of lysosome) could induce cell death in crizotinib-resistant NSCLC by inhibiting AA-promoted autophagy flux. Our study reveals a novel link between metabolic reprogramming, autophagy flux, and crizotinib resistance. We demonstrate that AA-driven autophagy flux contributes to resistance and propose CQ as a potential strategy to reverse this effect, providing new insights into crizotinib-resistant NSCLC therapy. We present this article in accordance with the MDAR reporting checklist (available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-2025-105/rc).
Methods
Cell culture and establishment of crizotinib-resistant cell lines
ROS1-rearranged HCC78 and ALK-rearranged H3122 NSCLC cell lines were obtained from the Cell Bank of Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). The cells were cultured in RPMI-1640 medium (HyClone, Logan, UT, USA) supplemented with 10% fetal bovine serum (FBS) (HyClone) and 100 units/mL of penicillin and 100 mg/L of streptomycin (HyClone) at 37 ℃ with 5% CO2. To establish crizotinib-resistant cell lines, HCC78 and H3122 cells were treated with incrementally increasing concentrations of crizotinib starting at 0.2 µM [below the half-maximal inhibitory concentration (IC50) of HCC78 and H3122 cells] (41). After 4 months, the cells that could grow in the presence of 2 or 1 µM of crizotinib were designated as HCC78CR and H3122CR cells, respectively. The HCC78CR and H3122CR cells were maintained in culture medium containing 1 or 0.5 µM of crizotinib, respectively.
Reagents
Crizotinib was purchased from Selleck Chemicals (Houston, TX, USA). AA and CQ were purchased from MedChemExpress (Monmouth Junction, NJ, USA).
Cell viability assay
The cell viability and IC50 of crizotinib were assessed using Cell Counting Kit-8 (CCK-8; Beyotime, Nanjing, China). Briefly, HCC78 or HCC78CR cells (1×103 cells/well) were placed in 96-well plates and treated with different concentrations of crizotinib (40, 20, 10, 5, 2.5, 1.25, 0.625, and 0.3125 µM). All experiments were performed with concentrations and time of exposure as indicated. After 48 hours, the culture media were replaced, and cells were incubated with 100 µL of CCK-8 solution for an additional 2 hours. Absorbance at 450 nm was measured using a microplate spectrophotometer (Agilent Technologies, Santa Clara, CA, USA). The IC50 value was determined as the crizotinib concentration that resulted in 50% cell death. Each assay was performed three or more times.
Colony formation assays
Cells (2×103) were seeded in each well of a six-well plate and cultured for 2 weeks. Forty-eight hours after inoculation, cells were treated with crizotinib. The medium was changed every 2–3 days, and cells were cultured with inhibitors for 9 days to 2 weeks. Colonies were fixed in 4% paraformaldehyde (PFA) (Solarbio, Beijing, China) for 15 minutes and stained with 1% crystal violet (KeyGen Biotech, Nanjing, China) for 20 minutes at room temperature. After staining, pictures of the wells were taken, and the number of colony-forming units was manually counted in three independent experiments.
Cell apoptosis analysis
An annexin V-APC assay kit (KeyGen Biotech) was used to detect apoptosis. Briefly, 1×106 HCC78/HCC78CR cells were seeded in each well of a six-well plate and cultured until 90% confluence. After trypsinization and suspension, the cells were stained with 2 µL annexin V-APC and 2 µL propidium iodide (PI) in 400 µL of annexin V-binding buffer, followed by incubation in the dark for 15 minutes. Flow cytometry was performed using a Cytomics FC 500 flow cytometer (Beckman Coulter, Brea, CA, USA), and data were analyzed using FlowJo software (BD Biosciences, Franklin Lakes, NJ, USA).
Autophagy flux assay
Cells were seeded in a Costar flat-bottom plate (cat. no. 3513; Corning, Corning, NY, USA) and transfected with a plasmid encoding the tandem fluorescent probe mCherry-GFP-LC3. After 24 hours, the cells were treated with 2 µM crizotinib or 30 µM CQ for 24 hours. Cells were fixed with 4% paraformaldehyde (cat no. P1110; Solarbio) for 15 minutes at room temperature. After being washed with phosphate-buffered saline (PBS), the cells were permeabilized with 0.3% Triton X-100 (cat. no. T8200; Solarbio) in PBS for 10 minutes and stained with DAPI (cat. no. C0060; Solarbio) for 5 minutes. The cells were then mounted for analysis and examined under a microscope (Leica Microsystems, Wetzlar, Germany). Cells with suitable mCherry-GFP-LC3 fluorescence were selected and quantified using the Analyze Particles plugin in ImageJ software.
Transmission electron microscopy analysis
HCC78 and HCC78CR cells were treated with or without 2 µM crizotinib for 24 hours and centrifuged at 1,000 ×g for 3 minutes after trypsin digestion, and cell precipitates were fixed with 4% glutaraldehyde at 4 ℃ for 24 hours. Subsequently, cells were fixed with 1% osmium tetroxide for 1 hour at 4 ℃. Following fixation, cells were dehydrated in a graded series of alcohol and acetone and then embedded in Epon 816 (Electron Microscopy Sciences, Hatfield, PA, USA). Ultrathin sections were obtained using a Leica ultramicrotome (Leica Microsystems) and then stained with uranyl acetate and lead citrate. Transmission electron microscopy images were captured with a JEM-1400 Plus transmission electron microscope (JEOL Ltd., Tokyo, Japan).
Measurement of reactive oxygen species (ROS) generation in cells
The intracellular ROS level was detected via flow cytometry. Briefly, 5×105 cells were placed in six-well plate and cultured overnight. Cells were then treated with 2µM crizotinib or/and 40µM AA for 24 hours. Following treatment, cells were stained with 10 µM of 2',7'-dichlorofluorescin diacetate (DCFH-DA) (Beyotime) at 37 ℃ for 30 minutes. After staining, cells were harvested and washed three times with ice-cold PBS, and their fluorescence was measured using a Cytomics FC 500 flow cytometer (Beckman Coulter). In all experiments, 10,000 viable cells were analyzed.
RNA isolation and gene expression analysis
RNA was extracted from cells and tissues using the Multisource Total RNA Mini Kit (Axygen Scientific, Union City, CA, USA). Subsequently, complementary DNA (cDNA) was synthesized from the extracted RNA using a PrimeScript RT reagent Kit with a gDNA Eraser Kit (cat. no. RR047Q; Takara Bio, Kusatsu, Japan) according to the manufacturer’s protocol. The synthesized cDNA served as the template and was mixed with TB Green Premix Ex Taq Kit (cat. no. RR420Q; Takara Bio) and target-specific primers; the sequences (5'-3') were as follows: CHOP-forward, GGAAACAGAGTGGTCATTCCC; CHOP-reverse, CTGCTTGAGCCGTTCATTCTC; sXBP1-forward, CTGAGTCCGAATCAGGTGCAG; sXBP1-reverse, ATCCATGGGGAGATGTTCTGG; PTGS2-forward, TCCCTTGGGTGTCAAAGGTAAA; PTGS2-reverse, CCTGGGGATCAGGGATGAAC; ACTB-forward, CATGTACGTTGCTATCCAGGC; and ACTB-reverse, CTCCTTAATGTCACGCACGAT.
Real time quantitative polymerase chain reaction (RT-qPCR) was performed using a CFX96 Fast Real-Time PCR system (Bio-Rad Laboratories, Hercules, CA, USA). ACTB was used as the internal control, and the relative expression level of each gene was calculated using the 2−ΔΔCt method. Triplicate assays were performed for each sample to ensure reliability. Solubility curves were analyzed to confirm the specificity of amplification products and exclude nonspecific amplification.
Western blotting and human phospho-kinase array
For experiments with protein analysis, cells were treated with specific chemical agents and collected via centrifugation. Cells were placed in RIPA buffer (Beyotime) containing protease inhibitors (Selleck Chemicals) and allowed to dissolve for 30 minutes on ice. The supernatant of the protein solution was collected via centrifugation at 17,000 ×g for 15 minutes at 4 ℃. Protein concentrations were determined using a Bicinchoninic acid Protein Assay Kit (cat no. 2312003; Solarbio). Cleared lysates were mixed with protein loading buffer (Beyotime) and boiled for 5 minutes. Samples were loaded onto sodium dodecyl sulfate-polyacrylamide gels, resolved by electrophoresis, and transferred to polyvinylidene fluoride membranes (MilliporeSigma, Burlington, MA, USA). The membranes were probed with the specified antibodies. Protein bands were visualized using a chemiluminescence imaging system (Clinx Science Instruments Co., Ltd., Shanghai, China) with chemiluminescent horseradish peroxidase (HRP) substrate and enhanced chemiluminescence (MilliporeSigma).
The primary antibodies used were LAMP1 [cat. no. 9091; Cell Signaling Technology (CST), Danvers, MA, USA], LC3B (cat. no. 83506; CST), p-p38MAPK (Thr180/Tyr182) (cat. no. 4511; CST), p38 MAPK (cat. no. 8690; CST), p-STAT3 (S727) (cat. no. 9134; CST), STAT3 (cat. no. 9139; CST), p-IRE1 (S724) (cat no. AP1442; ABclonal, Woburn, MA, USA), IRE1 (cat no. A17940; ABclonal), sXBP1 (cat no. A17007; ABclonal), p-PERK (T982) (cat no. AP0886; ABclonal), and PERK (cat. no. A18196; ABclonal). β-actin, HRP-conjugated goat anti-rabbit, and goat anti-mouse secondary antibodies were obtained from Boster Bio (Pleasanton, CA, USA).
In addition, a human phosphokinase array kit (cat no. ARY003C; R&D Systems; Minneapolis, MN, USA) was used to analyze the phosphorylation status of various kinases according to the manufacturer’s instructions. The array was incubated with cell lysates, and the phosphorylated proteins were detected using chemiluminescence.
RNA-sequencing analysis
Messenger RNA sequencing (mRNA-seq) and quality checks were conducted by HuaDa Gene Information Technology (Shenzhen, China). Briefly, HCC78 and HCC78CR cells (2×105 cells/well) were seeded into six-well plates. The cells were collected, and total RNA was extracted using TRIzol (Invitrogen, Thermo Fisher Scientific, Waltham, MA, USA). Three samples were collected in each group, and meaningful probe sets with a fold-change >1.5 and P value <0.05 (Student’s t-test) and a false-discovery rate (FDR) <0.05 were screened. Differentially expressed genes were then enriched via Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways.
Analysis of single-cell RNA sequencing data
The single-cell RNA sequencing (scRNA-seq) dataset, which provides transcriptional profiles of individual cells to uncover cellular heterogeneity, was obtained with accession code GSE223779 from the Gene Expression Omnibus (GEO) repository. This dataset includes both TKI-sensitive and TKI-resistant lung cancer organoid samples, derived from resected primary ALK-rearranged lung adenocarcinoma (42). The “Seurat” package in R version 4.0 (The R Foundation for Statistical Computing) was used for data processing. Quality control was performed in accordance with a previous study (42). Gene expression matrices were variance-stabilized using R package scTransform, a normalization method that models gene expression as a function of sequencing depth and uses regularized negative binomial regression to improve data quality and mitigate technical variability (43). Principal component analysis (PCA) was employed for dimensionality reduction of the scRNA-seq data. Uniform manifold approximation and projection (UMAP) was applied to visualize cell clusters. The Find Clusters function from R package Seurat was used to identify cell clusters. Oncogenic signaling pathway activity scores were calculated using the R package “progeny” (44), while trajectory analyses were performed using the R package “monocle2” (45).
Metabolomics profiling
Intracellular metabolites were analyzed using ultra high-performance liquid chromatography-mass spectrometer (UPLC-MS) (XevoG2-sQ-TOF, Waters). Briefly, 5×106 cells were lysed in ice-cold methanol and water (80:20). The extracted metabolites were dried, resuspended in methanol, and centrifuged at 13,000 ×g at 4 ℃ for 10 minutes. The supernatant was used for UPLC-MS analysis.
Metabolite identification and data analysis were performed using Progenesis QI software (Nonlinear Dynamics, Waters Corporation, Milford, MA, USA) and EZinfo software (Waters Corporation). Metabolites were identified with reference to the Lipid Maps Database (www.lipidmaps.org) and the Human Metabolome Database (http://www.hmdb.ca/). Metabolites with variable importance (VIP) >1 and P<0.05 between the HCC78 and HCC78CR were considered significantly different. Enrichment analysis of metabolic pathways was performed via the MetaboAnalyst website (https://www.metaboanalyst.ca).
Statistical analysis
Data were analyzed using GraphPad Prism version 9.0 (Graph Pad Software, Inc., Boston, MA, USA). In vitro data are presented as the mean ± SD. The two-sample t-test was used to determine significance with equal variance. The statistical significance of differences between more than two groups was assessed using one-way analysis of variance (ANOVA) or two-way ANOVA followed by the Dunnett multiple-comparison test. Two-sided P values less than 0.05 were considered significant.
Results
Crizotinib induced growth inhibition and apoptosis in sensitive cells compared to resistant cells
To clarify the mechanism of crizotinib resistance on NSCLC cells, we first established a model for crizotinib resistance in the HCC78 cells harboring the SLC34A2–ROS1 fusion and in the H3122 cells harboring the EML4–ALK fusion, by exposing them to increasing concentrations of the drug. After 8 weeks of crizotinib treatment, the IC50 significantly increased in crizotinib-resistant HCC78 (designated as HCC78CR) and H3122 (designated as H3122CR) cells compared to their corresponding parental cells (Figure 1A). After treating HCC78 cells with 1 µM and H3122 cells with 0.5 µM of crizotinib, we observed a time-dependent inhibition of cell proliferation compared to the corresponding resistant cells (Figure 1B). Colony formation assays showed that both HCC78CR and H3122CR cells increased the number of generated colonies under crizotinib treatment as compared to their respective parental cells (Figure 1C,1D).

To determine whether apoptosis is involved in crizotinib-induced cell death in sensitive cells, we examined apoptotic cell death using annexin V and PI co-staining followed by flow cytometry. The results confirmed that the apoptosis rates in HCC78CR and H3122CR cells induced by crizotinib were reduced compared to those in sensitive cells (Figure 1E,1F). Furthermore, as additional measurement of apoptosis, the levels of cleaved poly (ADP-ribose) polymerase (PARP) were higher in both HCC78 and H3122 cells following crizotinib treatment in a concentration-dependent manner as compared to resistant cells (Figure 1G,1H). Taken together, these data indicate that crizotinib induces growth inhibition and apoptosis more effectively in sensitive NSCLC cells compared to resistant cells. Notably, as indicated by the results shown in Figure 1, H3122CR cells exhibited less resistance to crizotinib than HCC78CR cells (lower IC50, fewer growing colonies, more crizotinib-induced apoptosis).
Crizotinib promoted the accumulation of autophagosomes by blocking autophagy flux in sensitive cells
Recent research suggests that CD74-ROS1 L2026M mutant contributes crizotinib resistance in NSCLC by stimulating autophagy (24). We therefore evaluated the potential effects of crizotinib on autophagy in NSCLC cells. Increased levels of microtubule-associated protein 1 light chain 3II (MAP1LC3II) can serve as an indicator of autophagy initiation and are commonly used as a marker for autophagosomes. Intriguingly, western blot analyses revealed that crizotinib treatment significantly increased the conversion of LC3I to LC3II and the protein expression of lysosomal-associated membrane protein 1 (LAMP1; a lysosome marker) in sensitive HCC78 and H3122 cells as compared to their resistant cells (Figure 2A,2B). Transmission electron microscopy analysis consistently indicated that autophagosomes were significantly accumulated in crizotinib-treated sensitive cells but not in resistant cells (Figure 2C). These results suggest that crizotinib promotes the accumulation of autophagosomes.

However, a greater abundance of autophagosomes does not necessarily imply a greater degree of autophagy (46). The significant rise in autophagosome levels can result from either the induction of autophagy or the impairment of autophagic flux. The protein level of the cargo sequestosome 1 [SQSTM1, also known as P62, a marker of impaired autophagic flux (47)] was higher in HCC78 cells following crizotinib treatment, but this increase was not observed in HCC78CR cells, while no significant change was observed in H3122/H3122CR cells (Figure 2A,2B). Additionally, HCC78 and HCC78CR cells were transfected with the mCherry-GFP-LC3 reporter plasmid, a specific indicator of autophagic vacuoles, and then treated with 2 µM of crizotinib. More mCherry-LC3 dots (red spots indicate autolysosomes) were observed in HCC78CR cells than in HCC78 cells (Figure 2D,2E), suggesting that resistant cells exhibited stronger autophagy activity than sensitive cells. After crizotinib treatment, the number of colocalized mCherry-LC3 dots and GFP-LC3 dots (yellow spots indicate autophagosomes) were significantly increased in HCC78 cells, but there was no significant change in HCC78CR cells (Figure 2D,2E). These results suggest that crizotinib blocks autophagic flux, leading to excessive autophagosome accumulation in sensitive NSCLC cells.
Crizotinib-induced accumulation of autophagosomes triggered ROS production and ER stress
Previous investigations have indicated that suppressing autophagy in pancreatic tumor cells leads to ROS production, heightened DNA damage, and impaired metabolic efficiency (48). Our results confirmed that crizotinib inhibits autophagy activity. The production of ROS in HCC78 and HCC78CR cells treated with crizotinib for 24 hours was measured using DCFH-DA staining. As shown in Figure 3A, 2 µM of crizotinib significantly promoted ROS upregulation in HCC78 cells as compared to resistant cells. Moreover, Western blotting showed that the level of p-H2A.X(S139), an indicator for DNA damage, was increased significantly in sensitive cells after crizotinib treatment compared to resistant cells (Figure 3B).
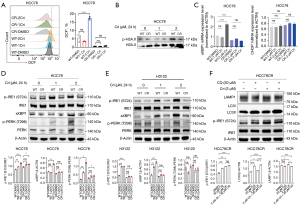
Elevated ROS production can lead to the accumulation of unfolded proteins in the ER, triggering an ER stress response. In this regard, the mRNA levels of ER effector proteins spliced X-box binding protein 1 (sXBP1) and DNA damage-inducible transcript 3 (CHOP) were measured using RT-qPCR. We found that 2 µM of crizotinib treatment significantly increased the expression of sXBP1, but not that of CHOP, in HCC78 cells (Figure 3C). We further evaluated the activity of ER transmembrane sensors, inositol-requiring enzyme1 (IRE1; also known as ERN1) and double-stranded RNA-activated protein kinase (PKR)-like ER kinase (PERK; also known as EIF2AK3) via western blotting. The results showed that p-IRE1 (S724) and sXBP1, but not p-PERK (T982), were upregulated by crizotinib in HCC78 and H3122 cells (Figure 3D,3E). These findings strongly suggest that crizotinib significantly induced IRE1/sXBP1-mediated ER stress in sensitive cells but not in resistant cells.
To investigate the regulation of autophagy on ER stress, CQ (an inhibitor of the lysosomal function) was used to inhibit autophagy in HCC78CR cells. The results showed that CQ significantly increased LC3II and promoted protein levels of p-IRE1 (S724) in HCC78CR cells (Figure 3F). Together, these data demonstrated that dysfunctional autophagosome accumulation induced by crizotinib promoted ROS levels and ER stress in NSCLC cells.
Downregulation of MAPK/p-STAT3/PTGS2 drove metabolic reprogramming in crizotinib-resistant cells
To investigate the mechanism of crizotinib resistance via the maintenance of autophagy activity in NSCLC cells, we first analyzed published scRNA-seq data from three ALK-rearranged lung cancer organoid samples, including one TKI-sensitive and two TKI-resistant samples from the same source (42). A total of 25,830 genes in 18,105 cells were included in the downstream analyses after quality control. UMAP analysis and visualization showed significant differences between TKI-sensitive and TKI-resistant cells (Figure 4A). Two TKI-sensitive clusters (SensC1 and SensC2) and six TKI-resistant clusters (ResiC1–ResiC6) were identified, indicating remarkable transcriptional heterogeneity within each treatment group (Figure 4A). Figure 4B presents the top 5 marker genes in each cluster, with cluster ResiC2, ResiC5, and ResiC6 sharing similar marker gene expression patterns. The oncogenic signaling pathway activity was also estimated in each cluster, revealing that the phosphoinositide 3-kinase (PI3K) signaling pathway was enhanced in most TKI-resistant clusters except ResiC1 (Figure 4C). Surprisingly, multiple signaling pathways, including JAK-STAT, MAPK, TNFα, and NF-κB, were significantly downregulated in TKI-resistant clusters (Figure 4C).

Further, a human phospho-kinase array analysis was performed to evaluate the signaling pathways in HCC78 and HCC78CR cells. The result showed that compared to sensitive cells, p-CREB (S133), p-GSK3α/β (S21/S9), p-WNK1(T60), p-YES (Y426), and p-P53(S46) were upregulated, while HSP60, p-RSK1/2(S221/S227), and p-STAT3(S727) were downregulated in HCC78CR cells treated with crizotinib for 24 hours (Figure 4D). Notably, p-STAT3 (S727) was downregulated in HCC78CR cells, which is consistent with the above-mentioned results regarding the downregulation of the JAK-STAT signaling pathway in most of TKI-resistant clusters of ALK-rearranged lung cancer organoids (Figure 4C). Moreover, we confirmed that both p-p38MAPK (Thr180/Tyr182) and p-STAT3 (S727) were downregulated in HCC78CR cells compared to parental cells via western blotting (Figure 4E). These data suggest that the MAPK/STAT3 signaling pathway was significantly inactivated after crizotinib resistance both in vitro and in vivo.
To identify the downstream target genes of transcription factor STAT3 involved in crizotinib resistance, RNA-seq analysis was performed to evaluate gene expression in HCC78 and HCC78CR cells. We found that 1240 genes were significantly upregulated [log2 fold change (FC) ≥2 and q<0.05] in HCC78 CR cells compared to HCC78 cells, while 483 genes were significantly downregulated (log2 FC ≤–2 and q<0.05) (Figure 4F). KEGG pathway enrichment analysis revealed that the differentially expressed genes were involved in IL-17 and cytokine–cytokine receptor interaction signaling pathway (Figure 4G). The IL-17 signaling pathway includes NF-κB and MAPK signaling pathway, which participate in signaling cascades controlling cellular responses to cytokines and stress, thereby regulating multiple transcription factors, including STAT3.
Furthermore, we analyzed the key node genes of four differential signaling pathways (lL-17 signaling pathway, cytokine-cytokine receptor interaction, NF-κB signaling pathway, and regulation of lipolysis in adipocytes) through computational network (CNET) mapping. We found that the metabolic enzyme PTGS2 was significantly downregulated and involved in all of these signaling pathways (Figure 4F,4H). In the aforementioned published scRNA-seq data, all clusters showed high expression of EPCAM, which indicates that the cells in the organoids are epithelial-derived tumor cells (Figure 4I). Correspondingly, PTGS2 expression was particularly attenuated in five TKI-resistant clusters, including ResiC1, ResiC2, ResiC3, ResiC4, and ResiC5 (Figure 4I). Additionally, we performed trajectory analyses to determine the differentiation trajectory of TKI-sensitive and TKI-resistant cells. Pseudotime analysis demonstrated a cell development trajectory from TKI-sensitive to TKI-resistant clusters (Figure 4J). Cluster ResiC2, ResiC5, and ResiC6 were primarily distributed at the terminal of the development trajectory, which aligned with the expression of PTGS2 (Figure 4J). RT-qPCR further confirmed that PTGS2 mRNA levels were reduced in HCC78CR and H3122CR cells as compared to their corresponding parental cells (Figure 4K). Collectively, these results suggest that the MAPK/p-STAT3-PTGS2 axis was downregulated and played a crucial role in dictating metabolic reprogramming in crizotinib-resistant NSCLC cells.
AA rescued the ROS level and ER stress induced by crizotinib via decreasing autophagosome accumulation
PTGS2 is a key enzyme in the conversion of AA to prostaglandins and other eicosanoids, and it was downregulated in crizotinib-resistant NSCLC cells compared to their corresponding parental cells. To investigate the mechanism of PTGS2 involvement in crizotinib resistance through metabolic programming, we performed a metabolomics assay by UPLC-MS analysis on HCC78CR cells and their corresponding parental cells. We identified 54 different metabolites between HCC78CR and HCC78 cells (Figure 5A). KEGG pathway enrichment analysis showed that the different metabolites were involved in the Warburg effect, glutathione metabolism, folate metabolism, malate-aspartate shuttle, AA metabolism and pentose phosphate pathway (Figure 5B). Accordingly, the abundance of polyunsaturated fatty-acid AA was significantly elevated, while metabolite prostaglandin E2 (PGE2) was decreased in HCC78CR cells as compared to HCC78 cells (Figure 5C). These results suggest that lipid composition after crizotinib resistance may be mediated by PTGS2.

Furthermore, the potential roles of AA in regulating autophagy activity were investigated. Interestingly, we found that AA significantly counteracted the elevation of LC3II and LAMP1 protein levels induced by crizotinib in HCC78 and H3122 cells, as shown by western blot analyses (Figure 5D). Moreover, the inhibition of mCherry-LC3 dots (red spots, representing the autolysosomes) by crizotinib was rescued by AA in HCC78 cells (Figure 5E,5F). Consistently, the number of colocalized mCherry-LC3 dots and GFP-LC3 dots (yellow spots, representing the autophagosomes) in HCC78 cells was significantly decreased by AA (Figure 5E,5F). These results suggest that AA could decrease the crizotinib-induced autophagosome accumulation by promoting autophagy flux. Moreover, AA counteracted crizotinib-induced ROS generation in HCC78 cells (Figure 5G), and ER stress was also reduced through a decrease in the protein levels of p-IRE1 (S724) and sXBP1 (Figure 5H). These findings indicate that AA can rescue ROS and ER stress by reducing autophagosome accumulation.
Targeting AA-modulated autophagy enhanced sensitivity of crizotinib therapy
To determine whether the regulation of autophagy flux by AA is responsible for crizotinib resistance, we first monitored the effect of different concentrations of AA on the viability of NSCLC cells. The results showed that low-dose AA (12.5 µM <AA <100 µM) promoted NSCLC cell survival, while high-dose AA (>100 µM) had the opposite effect (Figure 6A). Correspondingly, the cell growth inhibition induced by crizotinib was partially rescued by 40 µM of AA in HCC78 and H3122 cells (Figure 6B). These results confirm that AA contributes to crizotinib resistance in NSCLC cells. To determine whether autophagy is involved in AA-mediated crizotinib resistance in NSCLC cells, we blocked autophagy flux using 30 µM of CQ. The result showed that the inhibition of LC3II and LAMP1 accumulation by AA was significantly reversed by CQ in HCC78 and H3122 cells (Figure 6C). Interestingly, 30 µM of CQ blocked the restorative effect of AA on cell proliferation in NSCLC cells treated with crizotinib (Figure 6D). Additionally, CQ decreased mCherry-LC3 dots (red spots) and promoted colocalization of mCherry-LC3 dots and GFP-LC3 dots (yellow spots) in HCC78CR cells, as observed via confocal microscopy (Figure 6E,6F). Consistently, CQ also significantly sensitized resistant NSCLC cells to crizotinib in vitro (Figure 6G). Collectively, these results suggest that targeting AA-modulated autophagy with CQ could enhance the sensitivity of NSCLC cells to crizotinib therapy (Figure 6H).

Discussion
In this study, we found that autophagosome formation was significantly reduced in crizotinib-resistant cells compared to parental cells under drug stress and that blocking autophagy sensitized resistant cells to crizotinib. Mechanistically, the accumulation of AA, which was driven by the downregulation of p-STAT3 and PTGS2, reduced the autophagosome accumulation induced by crizotinib. This reduction in autophagosome accumulation contributed to the development of drug resistance. Preliminary data showed that modeling autophagy by CQ, known as an autophagic inhibitor, may enhance cancer treatments. CQ exhibits an antitumor effect by overcoming drug resistance to PI3K/AKT inhibitors in triple negative breast cancer models, and the combinations of CQ with ipatasertinib or taliselisib and chemotherapy was shown to potentiate the treatment efficacy by overcoming drug resistance (49). Furthermore, CQ has also demonstrated an increased radiosensitivity effect in human glioblastoma cell lines (50). Combined CQ with crizotinib in our study significantly inhibited AA-mediated crizotinib resistance in NSCLC. Despite some mechanisms of crizotinib resistance in ROS1- and ALK-rearranged NSCLC may be overcome by more potent TKIs or combinations of crizotinib with other TKIs or chemotherapy, other therapeutic strategies with different anchor point are needed. Therefore, we propose a novel role of impaired autophagy in reversing crizotinib resistance in NSCLC cells, which provides new insights into drug resistance.
Autophagy figures prominently in various physiological processes of cancer cells, including stress response, metabolism, drug resistance, and anticancer immunity (48), and it is being increasingly recognized for its involvement in tumorigenesis and resistance to cancer therapies (21). In our study, we found that crizotinib increased the accumulation of autophagosomes in NSCLC cells. However, it is important to note that an increase in autophagosome numbers does not necessarily indicate enhanced autophagy. Indeed, autophagosome accumulation often results from impaired trafficking to lysosomes without a concomitant change in autophagosome biogenesis, while an increase in autophagosome abundance may indicate reduced degradative activity (46). Consistently, we found that crizotinib impaired autophagy flux, leading to excessive accumulation of autophagosomes in ROS1+ or ALK+ NSCLC cells, a phenomenon that also occurs in cardiomyocytes and induces cardiotoxicity (51). These results suggest that crizotinib-induced autophagy damage may be a key mechanism of its cytotoxicity. In the future, we will continue to investigate the underlying mechanisms of crizotinib-induced autophagy damage.
Omega-6 PUFAs, including AA, are essential fatty acids that are critical to various biological functions. Notably, AA presents a paradoxical role in cancer development. Previous studies have shown that high doses of AA (100 µM) can induce apoptosis in a variety of tumor cells and exert a degree of antitumor activity (52,53). However, recent studies have shown that AA could maintain cell survival, inhibit apoptosis, and promote growth in colorectal cancer and ovarian cancer (54,55). Through a review of these studies on AA, we found that AA exhibits both pro- and antiapoptotic effects. This difference in outcomes may be attributed to the specific model or the concentration of AA employed in each study. In our study, we found that high-dose AA (>100 µM) inhibited the growth of NSCLC cells, whereas low-dose AA (12.5 µM <AA <100 µM) had an opposite effect.
In addition, previous studies have focused on the role of AA in promoting or inhibiting tumor cells via apoptosis and/or ER stress (33,36,56). However, the role and mechanism of AA in regulating TKI resistance via mediating autophagy remains unclear. In our study, we confirmed that AA modulated crizotinib resistance by promoting autophagy flux and reducing autophagosome abundance, thereby inhibiting the cell death induced by crizotinib treatment.
While our study provides valuable insights, there are several limitations of our study that might be addressed in future research. Firstly, our study is based on only one ALK+ and one ROS1+ NSCLC cell line. Therefore, it would be important in future studies to validate the involvement of AA accumulation in crizotinib resistance across additional NSCLC cell lines harboring ALK or ROS1 gene fusions. Moreover, the role of AA and its modulation of autophagy in crizotinib resistance should be validated in vivo using NSCLC tissue samples. Finally, it would be of interest to assess whether AA accumulation and autophagy modulation is also a mechanism of resistance to ALK- and ROS1-TKIs of new generation.
Conclusions
We found that the downregulation of PTGS2 and induced AA accumulation may be the necessary condition for crizotinib resistance in NSCLC cells via the regulation of autophagy. Targeting autophagic flux sensitizes resistant tumors to crizotinib. Our results identified a newly discovered role of AA in crizotinib resistance and potentially provides a novel treatment strategy for alleviating TKI resistance.
Acknowledgments
None.
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-2025-105/rc
Data Sharing Statement: Available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-2025-105/dss
Peer Review File: Available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-2025-105/prf
Funding: This work was supported by
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-2025-105/coif). E.M.U. has received research grants from AstraZeneca and Merck; speaker fees from Amgen; travel support related to participation in international scientific meeting from AstraZeneca, MSD, and Roche; payment for participation in Advisory Board from AstraZeneca and Pfizer. E.S.R. has received research grants from Sanofi and Takeda; reports honoraria for lectures from Amgen, AstraZeneca, Bayer, Boehringer Ingelheim, Bristol-Myers Squibb, Roche, Takeda; and received payment for participation in Advisory Board from Roche and Takeda. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Thai AA, Solomon BJ, Sequist LV, et al. Lung cancer. Lancet 2021;398:535-54. [Crossref] [PubMed]
- Shaw AT, Bauer TM, de Marinis F, et al. First-Line Lorlatinib or Crizotinib in Advanced ALK-Positive Lung Cancer. N Engl J Med 2020;383:2018-29. [Crossref] [PubMed]
- Shaw AT, Kim DW, Nakagawa K, et al. Crizotinib versus chemotherapy in advanced ALK-positive lung cancer. N Engl J Med 2013;368:2385-94. [Crossref] [PubMed]
- Shaw AT, Ou SH, Bang YJ, et al. Crizotinib in ROS1-rearranged non-small-cell lung cancer. N Engl J Med 2014;371:1963-71. [Crossref] [PubMed]
- Awad MM, Katayama R, McTigue M, et al. Acquired resistance to crizotinib from a mutation in CD74-ROS1. N Engl J Med 2013;368:2395-401. [Crossref] [PubMed]
- Song A, Kim TM, Kim DW, et al. Molecular Changes Associated with Acquired Resistance to Crizotinib in ROS1-Rearranged Non-Small Cell Lung Cancer. Clin Cancer Res 2015;21:2379-87. [Crossref] [PubMed]
- Roys A, Chang X, Liu Y, et al. Resistance mechanisms and potent-targeted therapies of ROS1-positive lung cancer. Cancer Chemother Pharmacol 2019;84:679-88. [Crossref] [PubMed]
- Drilon A, Somwar R, Wagner JP, et al. A Novel Crizotinib-Resistant Solvent-Front Mutation Responsive to Cabozantinib Therapy in a Patient with ROS1-Rearranged Lung Cancer. Clin Cancer Res 2016;22:2351-8. [Crossref] [PubMed]
- Gainor JF, Dardaei L, Yoda S, et al. Molecular Mechanisms of Resistance to First- and Second-Generation ALK Inhibitors in ALK-Rearranged Lung Cancer. Cancer Discov 2016;6:1118-33. [Crossref] [PubMed]
- Caumont C, Veillon R, Gros A, et al. Neuroendocrine phenotype as an acquired resistance mechanism in ALK-rearranged lung adenocarcinoma. Lung Cancer 2016;92:15-8. [Crossref] [PubMed]
- Lin JJ, Zhu VW, Yoda S, et al. Impact of EML4-ALK Variant on Resistance Mechanisms and Clinical Outcomes in ALK-Positive Lung Cancer. J Clin Oncol 2018;36:1199-206. [Crossref] [PubMed]
- Schneider JL, Lin JJ, Shaw AT. ALK-positive lung cancer: a moving target. Nat Cancer 2023;4:330-43. [Crossref] [PubMed]
- Ruzzi F, Angelicola S, Landuzzi L, et al. ADK-VR2, a cell line derived from a treatment-naïve patient with SDC4-ROS1 fusion-positive primarily crizotinib-resistant NSCLC: a novel preclinical model for new drug development of ROS1-rearranged NSCLC. Transl Lung Cancer Res 2022;11:2216-29. [Crossref] [PubMed]
- Perera RM, Stoykova S, Nicolay BN, et al. Transcriptional control of autophagy-lysosome function drives pancreatic cancer metabolism. Nature 2015;524:361-5. [Crossref] [PubMed]
- Poillet-Perez L, Xie X, Zhan L, et al. Autophagy maintains tumour growth through circulating arginine. Nature 2018;563:569-73. [Crossref] [PubMed]
- Ni B, Li Q, Zhuang C, et al. The nerve-tumour regulatory axis GDNF-GFRA1 promotes tumour dormancy, imatinib resistance and local recurrence of gastrointestinal stromal tumours by achieving autophagic flux. Cancer Lett 2022;535:215639. [Crossref] [PubMed]
- Yu T, Guo F, Yu Y, et al. Fusobacterium nucleatum Promotes Chemoresistance to Colorectal Cancer by Modulating Autophagy. Cell 2017;170:548-563.e16. [Crossref] [PubMed]
- Meng J, Liu K, Shao Y, et al. ID1 confers cancer cell chemoresistance through STAT3/ATF6-mediated induction of autophagy. Cell Death Dis 2020;11:137. [Crossref] [PubMed]
- Jiang GM, Tan Y, Wang H, et al. The relationship between autophagy and the immune system and its applications for tumor immunotherapy. Mol Cancer 2019;18:17. [Crossref] [PubMed]
- Papademetrio DL, Cavaliere V, Simunovich T, et al. Interplay between autophagy and apoptosis in pancreatic tumors in response to gemcitabine. Target Oncol 2014;9:123-34. [Crossref] [PubMed]
- Bhardwaj M, Leli NM, Koumenis C, et al. Regulation of autophagy by canonical and non-canonical ER stress responses. Semin Cancer Biol 2020;66:116-28. [Crossref] [PubMed]
- Cheng X, Feng H, Wu H, et al. Targeting autophagy enhances apatinib-induced apoptosis via endoplasmic reticulum stress for human colorectal cancer. Cancer Lett 2018;431:105-14. [Crossref] [PubMed]
- Wang Y, Zhou Q, Liu C, et al. Targeting IL-6/STAT3 signaling abrogates EGFR-TKI resistance through inhibiting Beclin-1 dependent autophagy in HNSCC. Cancer Lett 2024;586:216612. [Crossref] [PubMed]
- Xu X, Li Y, Xu R, et al. CD74-ROS1 L2026M mutant enhances autophagy through the MEK/ERK pathway to promote invasion, metastasis and crizotinib resistance in non-small cell lung cancer cells. FEBS J 2024;291:1199-219. [Crossref] [PubMed]
- Valencia K, Echepare M, Teijeira Á, et al. DSTYK inhibition increases the sensitivity of lung cancer cells to T cell-mediated cytotoxicity. J Exp Med 2022;219:e20220726. [Crossref] [PubMed]
- Sun L, Zhang H, Gao P. Metabolic reprogramming and epigenetic modifications on the path to cancer. Protein Cell 2022;13:877-919. [Crossref] [PubMed]
- Sounni NE, Cimino J, Blacher S, et al. Blocking lipid synthesis overcomes tumor regrowth and metastasis after antiangiogenic therapy withdrawal. Cell Metab 2014;20:280-94. [Crossref] [PubMed]
- Zhao Y, Butler EB, Tan M. Targeting cellular metabolism to improve cancer therapeutics. Cell Death Dis 2013;4:e532. [Crossref] [PubMed]
- Talebi A, Dehairs J, Rambow F, et al. Sustained SREBP-1-dependent lipogenesis as a key mediator of resistance to BRAF-targeted therapy. Nat Commun 2018;9:2500. [Crossref] [PubMed]
- Iwamoto H, Abe M, Yang Y, et al. Cancer Lipid Metabolism Confers Antiangiogenic Drug Resistance. Cell Metab 2018;28:104-117.e5. [Crossref] [PubMed]
- Yan C, Wu D, Gan L, et al. Significant metabolic alterations in non-small cell lung cancer patients by epidermal growth factor receptor-targeted therapy and PD-1/PD-L1 immunotherapy. Front Pharmacol 2022;13:949745. [Crossref] [PubMed]
- Martin SA, Brash AR, Murphy RC. The discovery and early structural studies of arachidonic acid. J Lipid Res 2016;57:1126-32. [Crossref] [PubMed]
- Wang B, Wu L, Chen J, et al. Metabolism pathways of arachidonic acids: mechanisms and potential therapeutic targets. Signal Transduct Target Ther 2021;6:94. [Crossref] [PubMed]
- Badimon L, Vilahur G, Rocca B, et al. The key contribution of platelet and vascular arachidonic acid metabolism to the pathophysiology of atherothrombosis. Cardiovasc Res 2021;117:2001-15. [Crossref] [PubMed]
- He S, Lu G, Hou H, et al. Saikosaponin-d suppresses the expression of cyclooxygenase-2 through the phospho-signal transducer and activator of transcription 3/hypoxia-inducible factor-1α pathway in hepatocellular carcinoma cells. Mol Med Rep 2014;10:2556-62. [Crossref] [PubMed]
- Bae S, Kim MK, Kim HS, et al. Arachidonic acid induces ER stress and apoptosis in HT-29 human colon cancer cells. Anim Cells Syst (Seoul) 2020;24:260-6. [Crossref] [PubMed]
- Li J, Li O, Kan M, et al. Berberine induces apoptosis by suppressing the arachidonic acid metabolic pathway in hepatocellular carcinoma. Mol Med Rep 2015;12:4572-7. [Crossref] [PubMed]
- Montecillo-Aguado M, Tirado-Rodriguez B, Huerta-Yepez S. The Involvement of Polyunsaturated Fatty Acids in Apoptosis Mechanisms and Their Implications in Cancer. Int J Mol Sci 2023;24:11691. [Crossref] [PubMed]
- Hu Y, Luo NJ, Gan L, et al. Heat stress upregulates arachidonic acid to trigger autophagy in sertoli cells via dysfunctional mitochondrial respiratory chain function. J Transl Med 2024;22:501. [Crossref] [PubMed]
- Dall KB, Færgeman NJ. Metabolic regulation of lifespan from a C. elegans perspective. Genes Nutr 2019;14:25. [Crossref] [PubMed]
- Kato Y, Ninomiya K, Ohashi K, et al. Combined effect of cabozantinib and gefitinib in crizotinib-resistant lung tumors harboring ROS1 fusions. Cancer Sci. 2018;109:3149-58. [Crossref] [PubMed]
- Kwok HH, Li H, Yang J, et al. Single-cell transcriptomic analysis uncovers intratumoral heterogeneity and drug-tolerant persister in ALK-rearranged lung adenocarcinoma. Cancer Commun (Lond) 2023;43:951-5. [Crossref] [PubMed]
- Hafemeister C, Satija R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol 2019;20:296. [Crossref] [PubMed]
- Schubert M, Klinger B, Klünemann M, et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat Commun 2018;9:20. [Crossref] [PubMed]
- Qiu X, Mao Q, Tang Y, et al. Reversed graph embedding resolves complex single-cell trajectories. Nat Methods 2017;14:979-82. [Crossref] [PubMed]
- Klionsky DJ, Abdalla FC, Abeliovich H, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012;8:445-544. [Crossref] [PubMed]
- Wang X, de Carvalho Ribeiro M, Iracheta-Vellve A, et al. Macrophage-Specific Hypoxia-Inducible Factor-1α Contributes to Impaired Autophagic Flux in Nonalcoholic Steatohepatitis. Hepatology 2019;69:545-63. [Crossref] [PubMed]
- Yang S, Wang X, Contino G, et al. Pancreatic cancers require autophagy for tumor growth. Genes Dev 2011;25:717-29. [Crossref] [PubMed]
- Cocco S, Leone A, Roca MS, et al. Inhibition of autophagy by chloroquine prevents resistance to PI3K/AKT inhibitors and potentiates their antitumor effect in combination with paclitaxel in triple negative breast cancer models. J Transl Med 2022;20:290. [Crossref] [PubMed]
- Ye H, Chen M, Cao F, et al. Chloroquine, an autophagy inhibitor, potentiates the radiosensitivity of glioma initiating cells by inhibiting autophagy and activating apoptosis. BMC Neurol 2016;16:178. [Crossref] [PubMed]
- Xu Z, Pan Z, Jin Y, et al. Inhibition of PRKAA/AMPK (Ser485/491) phosphorylation by crizotinib induces cardiotoxicity via perturbing autophagosome-lysosome fusion. Autophagy 2024;20:416-36. [Crossref] [PubMed]
- Vento R, D'Alessandro N, Giuliano M, et al. Induction of apoptosis by arachidonic acid in human retinoblastoma Y79 cells: involvement of oxidative stress. Exp Eye Res 2000;70:503-17. [Crossref] [PubMed]
- Serini S, Piccioni E, Merendino N, et al. Dietary polyunsaturated fatty acids as inducers of apoptosis: implications for cancer. Apoptosis 2009;14:135-52. [Crossref] [PubMed]
- Xu C, Gu L, Hu L, et al. FADS1-arachidonic acid axis enhances arachidonic acid metabolism by altering intestinal microecology in colorectal cancer. Nat Commun 2023;14:2042. [Crossref] [PubMed]
- Yang J, Zaman MM, Vlasakov I, et al. Adipocytes promote ovarian cancer chemoresistance. Sci Rep 2019;9:13316. [Crossref] [PubMed]
- Jia Z, Wu J, Liu F, et al. Arachidonic acid is involved in high-salt diet-induced coronary remodeling through stimulation of the IRE1α/XBP1s/RUNX2/OPN signaling cascade. Lipids Health Dis 2025;24:44. [Crossref] [PubMed]


