
Clinical application of minimal residual disease detection by ctDNA testing in non-small cell lung cancer: a narrative review
Introduction
According to the China Cancer Statistics 2022 (1), lung cancer is the leading cause of cancer-related deaths, accounting for approximately 20% of global cancer deaths. Non-small cell lung cancer (NSCLC) accounts for approximately 85% of lung cancer (2). Surgery is the main approach for patients with stage I-III NSCLC, however, the recurrence rate of the disease remains high (3,4). Imaging monitoring is a routine method for detecting tumor recurrence, but its sensitivity is not high, and effective means to evaluate treatment effectiveness are lacking, resulting in overtreatment or late intervention and inadequate treatment, which affects patient prognosis (5,6). Conventional protein markers cannot be used to monitor tumor recurrence because of their low sensitivity and specificity (7-9). Therefore, the timely and effective identification of patients with tumor recurrence is necessary in clinical practice to evaluate the treatment efficacy and accurately guide personalized treatment for patients with NSCLC.
Ultra-deep detection of circulating tumor DNA (ctDNA) in peripheral blood enables monitoring of minimal residual disease (MRD), which is not visible to the naked eye and involves tumor-derived gene mutations that can serve as prognostic biomarkers for patients with NSCLC (10-12). In this review, we evaluate the clinical significance of ctDNA in detecting MRD in patients with NSCLC, and address the challenges encountered during its implementation in clinical practice. We present this article in accordance with the Narrative Review reporting checklist (available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-24-942/rc).
Methods
We conducted a comprehensive search for published English articles in the PubMed and Web of Science databases from January 1, 2005, to September 14, 2024, utilizing keywords “non-small cell lung cancer” or “NSCLC” or “lung adenocarcinoma” or “LUAD” or “lung squamous carcinoma” or “LUSC” and “liquid biopsy” or “circulating tumor DNA” or “ctDNA” or “minimal residual disease” or “molecular residual disease” or “MRD”. We also searched the references of the included studies to supplement the acquisition of relevant information. Table 1 provides a comprehensive overview of the search strategies employed.
Table 1
| Items | Specification |
|---|---|
| Date of search | September 14, 2024 |
| Databases and other sources searched | PubMed; Web of Science |
| Search terms used | Non-small cell lung cancer or NSCLC or lung adenocarcinoma or LUAD or lung squamous carcinoma or LUSC and liquid biopsy or circulating tumor DNA or ctDNA or minimal residual disease or molecular residual disease or MRD |
| Timeframe | From January 2005 to September 2024 |
| Inclusion and exclusion criteria | Inclusion criteria: English-language articles; clinical research trials; translational clinical research |
| Exclusion criteria: non-English-language articles; clinical case report; abstracts; guidelines | |
| Selection process | Study selection were assessed by all authors |
ctDNA, circulating tumor DNA; LUAD, lung adenocarcinoma; LUSC, lung squamous carcinoma; MRD, minimal residual disease; NSCLC, non-small cell lung cancer.
MRD and ctDNA
The concept of MRD has been gradually extended from hematologic malignancies (such as acute lymphoblastic leukemia, multiple myeloma, and acute myeloid leukemia) to solid tumors (13-15). MRD is defined as the complete response (CR) to the disease achieved through surgery, radical radiotherapy, targeted therapy, or combination therapies, with a reduction in the tumor burden by >99%, and still residual tumor cells or other tumor cell derivatives, it can cause tumor recurrence (16-18). ctDNA is fragmented DNA derived from tumor cells, with a length of 130–150 base pairs and a half-life of 16 min to 2.5 h (19). It accounts for less than 1% of the nucleic acid fragments (cell-free DNA, cfDNA) (150–200 base pairs) released during cells apoptosis or necrosis in the blood (20,21). ctDNA contains information on genes related to about mutations, deletions, insertions, rearrangements, copy number abnormalities, and methylation. Thus, ctDNA detection can identify tumor-derived somatic mutations in the plasma, thereby identifying tumor recurrence (22-24).
Clinical value of ctDNA MRD in NSCLC
Identifying patients at high risk of recurrence
In 2017, a retrospective study by Chaudhuri et al. (25) demonstrated that in early-stage NSCLC, ctDNA MRD identified disease recurrence up to 5.2 months earlier than radiographic imaging, opening up therapeutic opportunities for patients with a low tumor burden. Another prospective study by Abbosh et al. (26) found that tracking NSCLC subclones by detecting ctDNA could predict recurrence and metastasis, with MRD positivity in 13 out of 14 patients with recurrence [detection of at least two single nucleotide variants (SNVs) represented MRD positivity], and recognition of disease recurrence a median of 70 days earlier (10–346 days). In 2020, Abbosh et al. (27) published the results from a study in which MRD was detected before or at the time of recurrence in 37 of 45 patients, predicting disease recurrence a median of 151 days (0–984 days) earlier than computed tomography (CT). ctDNA MRD is a promising biomarker for monitoring the postoperative recurrence of lung cancer (25,26).
In addition, Ohara et al. and Kuang et al. (28,29) found that preoperative ctDNA status was correlated with tumor size, and not independently associated with recurrence-free survival (RFS) (P=0.132 and P=0.61, respectively). However, Isaksson et al. found that preoperative ctDNA positivity was associated with early recurrence of lung adenocarcinoma (LUAD), but not with lung squamous cell carcinoma (LUSC), which may reflect biological differences between the two subtypes of NSCLC, such as significant upregulation of genes related to proliferation and cell cycle, and an elevated weighted genome integrity index in LUAD, leading to excessive tumor growth (27,30,31). Notably, the detection rate of preoperative ctDNA positive LUSC patients was higher than that of LUAD (97% vs. 19%), which may be due to the fact that LUSC patients are more susceptible to tumor cell necrosis and detachment (26,32). In addition to this, Peng et al. and Li et al. (33,34) found that ctDNA positivity was the main reason for RFS and overall survival (OS), regardless of whether ctDNA was detected pre or postoperatively. Compared with preoperative ctDNA-negative patients, ctDNA-positive patients have a 3.4- and 4-fold increased risk of recurrence and death, respectively (33). Therefore, compared with postoperative ctDNA status, the relationship between preoperative ctDNA status and patient prognosis has yielded inconsistent conclusions across various studies. This variability may be attributed to differences in sample size, histological subtypes, positive judgment criteria, and treatment methods (28,30,33). (I) Sample size is a critical factor that cannot be overlooked. Smaller sample sizes can increase the randomness of research outcomes, thereby failing to accurately reflect the true overall situation; (II) differences in histological subtypes are also a key determinant of research conclusions. Various subtypes of NSCLC exhibit significant variations in biological characteristics, malignancy, and metastatic potential; (III) inconsistencies in positive judgment criteria can lead to discrepancies in research results. Overly stringent criteria may misclassify patients with poor prognosis as negative, while overly lenient criteria may introduce an excessive number of false positives; (IV) the diversity of treatment methods significantly influences the relationship between preoperative ctDNA status and prognosis. Different treatment regimens, including surgical techniques, chemotherapy drug selection and dosages, and radiotherapy timing and dosages, can have substantial impacts on tumor progression and patient outcomes.
In summary, the varying conclusions regarding the relationship between preoperative ctDNA status and patient prognosis across different studies present a complex issue influenced by multiple factors. To more accurately elucidate this relationship, future research should focus on several key areas: first, conducting large-scale, multi-center studies to enhance sample size and improve the reliability and generalizability of findings; second, performing detailed classification and stratified analysis of patient types, taking into account factors such as tumor type and individual patient characteristics; third, standardizing predictive indicators and positive criteria to ensure comparability across studies; and fourth, considering the consistency of treatment methods in research design or conducting separate analyses for different treatment approaches.
Guiding adjuvant therapy
Treating patients with NSCLC with adjuvant therapy in clinical practice is mainly based on tumor-node-metastasis (TNM) staging; however, whether it is stage I or II–IIIA, some patients experience disease recurrence, which indicates the existence of hidden and latent MRD (35). Zhang et al. (36) retrospectively collected plasma samples from 73 patients after surgery and adjuvant chemotherapy (ACT), found that MRD-positive patients who received ACT were able to achieve a longer RFS than those who did not received ACT [median RFS 18.5 vs. 12.3 months, hazard ratio (HR) =0.63], while MRD-negative patients who did not receive ACT achieved a longer RFS than those who did receive ACT (median RFS 32.2 vs. 64.8 months, HR =0.09, P=0.028). This result was further validated in a prospective multicentre study with a large sample size by Xia et al. (31), which showed that compared with patients who were positive for ctDNA MRD at 3 days or 1 month post-surgery and received adjuvant therapy, all patients who did not receive adjuvant therapy relapsed, with a median RFS of 574 vs. 315 days (P=0.008). In contrast, MRD-negative patients who received adjuvant therapy had a recurrence rate of 27.7% (31/112), whereas patients who did not receive adjuvant therapy had a recurrence rate of 9% (18/191) (RFS was not achieved in either group; P<0.001). MRD-positive patients receiving adjuvant therapy may achieve longer RFS, whereas MRD-negative NSCLC patients are less likely to benefit from adjuvant therapy (37).
Dynamic monitoring of MRD changes may aid in identifying treatment-resistant populations. Abbosh et al. (26) found that ctDNA profiles responded to ACT resistance; three patients who were ctDNA-positive and received ACT had elevated ctDNA concentrations after treatment, and all relapsed within 1 year. In contrast, one patient had ctDNA turned negative during and after chemotherapy and remained disease-free for 688 days. Zhang et al. (38) found that patients with MRD turned negative after treatment were able to achieve a longer median disease-free survival (DFS) (14 vs. 16.3 vs. 22.5 months) compared to patients who remained MRD-positive after adjuvant therapy and transiently cleared the subsequently elevated MRD. Therefore, dynamic monitoring of MRD during treatment is helpful in determining therapeutic efficacy, as patients with elevated or persistently positive MRD levels do not respond to treatment, and such patients are advised to further tailor a precise and intensive treatment regimen. In contrast, patients with reduced or negative MRD responded well to treatment.
In conclusion, the ctDNA MRD status has the potential to predict therapeutic efficacy, and the MRD status after treatment may provide personalized treatment plans for patients with lung cancer (Figure 1). However, MRD sensitivity has been much lower than expected. In 2024, Bossé et al. (39) reported that ctDNA was detected in only 63 of 174 patients who relapsed, with a detection rate of 36%. In the same year, the ADAURA study reported at the American Society of Clinical Oncology (ASCO) showed that MRD detection guided the overall DFS sensitivity of the postoperative EGFR-mutated population targeted therapy to 65%, of which the DFS sensitivity of the placebo group was 70.8%. This means that if MRD testing is used to guide postoperative adjuvant therapy, at least 30% of patients with relapse will not receive adjuvant therapy, which undoubtedly reduces the number of people who benefit from adjuvant therapy. Therefore, ctDNA MRD should be combined with routine imaging monitoring to make a personalized treatment plan for each patient, and should not be too dependent on ctDNA MRD detection (40).
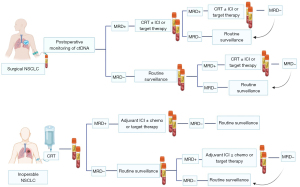
Clinical significance of ctDNA MRD in inoperable NSCLC
For patients with locally advanced and advanced inoperable NSCLC, local therapy combined with systemic therapy is the main treatment option; however, it often affects patients’ quality of life due to intolerable toxicity. Meanwhile, some patients with locally advanced disease are stable after chemoradiotherapy, with a 5-year progression-free survival (PFS) of 19%, and may be a potentially curable population (41,42). Similarly, data from clinical studies and real-world evidence showed that 28–46% of patients with advanced NSCLC had no disease progression within 3 years of treatment (43,44). Is de-escalation in potentially curable populations or those undergoing long-term stable treatment? Moding et al. conducted a retrospective study on this issue (45), exploring for the first time whether ctDNA MRD can guide the decision of implementing consolidation immune checkpoint inhibitor (ICI) therapy in patients with locally advanced NSCLC after chemoradiotherapy. The results showed that ctDNA MRD-negative patients did not benefit from consolidation therapy after receiving chemoradiotherapy, whereas ctDNA MRD-positive patients did benefit. Responders and non-responders were distinguished in the early stages of consolidation ICI treatment. Similarly, a recent prospective study (46) found that patients with early clearance of MRD during radiotherapy (n=44) had better PFS than those with late clearance (n=17) (22.9 vs. 10.3 months, P<0.001) and that patients with early clearance of MRD did not further benefit from consolidation therapy (P=0.917). The above two NSCLC data points indicate that ctDNA analysis may provide personalized solutions for whether local advanced stage patients receive consolidation therapy. Dong et al. conducted an exploratory proof-of-concept study to explore for the first whether ctDNA can guide de-escalation treatment in advanced-stage patients who have received complete response with targeted combined local therapy (47). If the imaging examination shows CR, and the patient has a plasma carcinoembryonic antigen and ctDNA-negative status, targeted therapy may be discontinued. If any of the results are abnormal during the discontinuation period, treatment will be restarted. Sixty patients were enrolled in the study, with a median follow-up time of 19.2 months, median PFS of 18.4 months, and a median total discontinuation time of 9.1 months. Among them, 14 patients were in the stage of drug discontinuation, during which the tumor did not recur. The median duration of drug discontinuation was 20.3 months, and the median PFS was not reached. Forty-six patients restarted targeted therapy owing to abnormal test results, which was safe and effective, with an efficacy rate as high as 96%, and half of the patients discontinued the therapy again because of the sufficient tumor regression. This indicates that de-escalation strategies are feasible for patients with CR and ctDNA-negative disease after receiving targeted and local therapy. A prospective translational study conducted by Xu et al. established an immune system subtypes (blood-based genomic immune subtypes, bGIS) scheme based on ctDNA integrated genomic features (48), dividing patients into three groups: bGIS-1 (ctDNA−), bGIS-2 [ctDNA+, blood-based tumor mutational burden (bTMB) high, blood-based intra-tumor heterogeneity (bITH) low, and blood-based chromosomal instability (bCIN) low], and bGIS-3 (bTMB low, bITH-high, bCIN-high). To explore whether the application of the bGIS method can identify ICI beneficiary population of patients with advanced NSCLC. The results suggested that bGIS-1 and bGIS-3 patients did not benefit from programmed cell death protein 1 (PD-1) inhibitors, whereas bGIS-2 patients did benefit from PD-1 inhibitors, which may indicate that developing refined stratification strategies for NSCLC patients based on bGIS approach is feasible.
In summary, recent clinical studies have provided novel treatment strategies for NSCLC patients, that is, ctDNA guided de-escalation therapy for patients with locally advanced and advanced stage NSCLC, and have developed a refined stratification for patients in need of treatment. However, the results of the present study should be interpreted with caution, and the clinical application of this new strategy requires more clinical research and more evidence-based medical evidence support.
Challenges in ctDNA detection of MRD
Sensitivity and specificity
In the field of solid tumors, current ctDNA detection technologies are primarily categorized into polymerase chain reaction (PCR) and next-generation sequencing (NGS). Owing to its high throughput, comprehensive coverage, and superior sensitivity, NGS-based MRD detection has emerged as the predominant approach. Nevertheless, NGS technology employed for MRD detection in early-stage NSCLC confronts two significant challenges.
On the one hand, the variant allele fractions (VAFs) detected in plasma reflect the amount of ctDNA released, which is usually proportional to the tumor volume. In the TRACERx study (26), the authors correlated nodule size obtained from low-dose CT scans with clonal mutations detected by plasma VAF, predicting that a primary tumor size of 10 cm3 (equivalent to T1c, 2.6 cm) corresponds to 0.1% of plasma VAF, and then further extrapolated to estimate that 1 cm3 (equivalent to T1a, 1.2 cm) tumor is equivalent to tumor plasma 0.008% [95% confidence interval (CI): 0.002–0.03%] of the mutant allele frequency (MAF) clonal mutation (49), However, existing detection techniques struggle to detect extremely low concentrations of ctDNA. For example, the CAncer Personalized Profiling by Deep Sequencing (CAPP-Seq) developed by Newman et al., a representative technology of tumor-naive assays, does not require prior information about tumor gene variants, and targets recurrently mutated 139 genes in lung cancer based on a fixed panel (50). The sequencing segment is condensed to 0.004% of the entire genome size, enabling ultra-deep sequencing to accurately detect ctDNA mutations with MAF as low as 0.02%. However, this assay has limitations, as 50% of the genomic positions have errors when the MAF was <0.02%. Therefore, to minimize errors, the researchers used integrated digital error suppression methods (iDES) (51), which combine unique identifiers (UIDs) and background polishing, to increase sensitivity while reducing errors by a factor of 15 (<1.5×10−5 errors per base), with a detection limit as low as 0.004%. Phallen et al. also developed a targeted error correction sequencing method (TEC-Seq) (52), which reduces sequencing errors to <3×10−7 per base of false-positive mutations through high-depth sequencing, optimized library generation, and the introduction of UIDs. These data indicate that the application of the current technology to detect low-frequency mutations may be limited by technical errors, resulting in false-positive test results. Therefore, achieving ultra-deep sequencing while reducing error rates may be a direction for future detection.
On the other hand, concentration of ctDNA in the early stages of NSCLC is low and lacks sensitivity. In the TRACERx study (26), Signatera technology used tumor tissue for whole exome sequencing (WES), customizing personalized panel for each patient to detect SNVs to identify tumor recurrence and metastasis, although the detection limit was as low as 0.01% and the specificity was >99.6%. However, the detection rate of early lung adenocarcinoma was only 19%. The updated results of the 2020 TRACERx series of studies showed that ArcherDx technology improved detection sensitivity by adding unique molecular identifiers (UMI) to the Signatera technology, with a 49% detection rate of early stage lung adenocarcinoma and a low detection limit of 0.003% (26,27). Technologies such as MRDetect, MRD-EDGE, and PhasED-Seq use the sequencing width to detect tumor mutations. MRDetect technology (53) detects SNVs and copy number alterations from whole genome sequencing of matched tumor tissue and peripheral blood mononuclear cells (PBMC) germline, achieving detection of ctDNA at a low sequencing depth (35X) and with a detection of ctDNA at tumor fraction (TF) sensitivities as low as 10−5, with a detection sensitivity of up to 67% in early stage LUAD, compared to 19% with Signatera technology (26). Based on the MRDetect platform, the research team made further breakthroughs and developed a plasma whole-genome sequencing ctDNA signal enrichment platform guided by advanced machine learning strategies “MRD-EDGE”. This platform can monitor tumor burden with high sensitivity and identify postoperative MRD in multiple types of tumors. When the TF is 5×10−5, the detection sensitivity can reach 0.7, achieving dynamic tracking of plasma TF after neoadjuvant ICI treatment in NSCLC, as well as the detection of postoperative MRD in other types of tumors (54). Based on this, PhasED-Seq technology is an alternative method of duplex sequencing to detect multiple mutations (phased variants, PVs) in the same DNA fragment, which reduces the error rate and increases the sensitivity of detection (55). In 24 plasma samples from six patients with solid tumors (five lung cancer and one breast cancer), PhasED-Seq detected ctDNA from six other plasma samples, in addition to CAPP-Seq based on nine samples, with a TF as low as 0.000094%.
Several of these techniques are representative of tumor-informed assays that require genetic testing of the primary tumor tissue and customization of a personalized regimen, an approach that increases sensitivity and specificity. However, owing to the heterogeneity among research cohorts, direct comparisons of the sensitivity and specificity across different MRD detection methods present a substantial challenge. Benchmarking different assays within the same patient cohort would be highly beneficial for addressing this issue, but it is necessary to ensure that sufficient blood volume can be obtained. According to previous studies (25,45,56), the median plasma cfDNA concentration after completing local therapy ranged from 3.0 to 10.8 ng/mL, which is equivalent to 900–3,300 haploid genomes per mL of plasma, with one haploid human genome having a mass of ~3.3 pg. In the MEDAL study published in 2023 (57), a head-to-head comparison of fixed and personalized MRD assays (the PROPHET method based on a tumor tissue WES assay containing 50 highly ranked variants) revealed that the sensitivity of personalized customized MRD panels was significantly higher than that of fixed MRD detection (42% vs. 17%), with a lower limit of detection (LOD) as low as 0.004%. In 2023, the European Society for Medical Oncology (ESMO) reported the findings of Gao et al., which conducted a head-to-head comparison between the personalized MRD detection method MNavifator V2, based on comprehensive genomic profiling, and the fixed-panel-based MNavifator V1. The study demonstrated that MNavifator V2 could detect NSCLC recurrence 15 days earlier than MNavifator V1, with a LOD as low as 0.005% (58). This result may further substantiate the superior sensitivity of tumor-informed assays compared to tumor-naive assays. However, the disadvantages of tumor-informed assays include the difficulty in obtaining tumor tissue, the inability to detect new tumors (based on the design of primary tumor-specific mutations), high cost of detection, and long detection time, which limit their clinical application. In contrast, compared to tumor-informed assays, tumor-naive assays provide comprehensive coverage of hotspot variants and have low detection costs and short times, but relatively low sensitivity and specificity. Therefore, exploring techniques with high sensitivity and low cost is necessary (Table 2).
Table 2
| Study | Platform | LOD | Sensitivity | Specificity | Application | Technology | Panel |
|---|---|---|---|---|---|---|---|
| Cohen 2018, (4) | CancerSEEK | – | 70% | >99% | Earlier detection cancer and localized the identify the anatomical location of the primary tumor | Tumor-naive assays | 61-amplicons [include 16 genes (4.6 kbp)] |
| Abbosh 2017, (26) | Signatera | 0.01% | 84% | 99.6% | MRD detection and validation of phylogenetic characterization | Personalizedtumor-informed assays | Median of 18 patient-specific SNVs (~1.5 kbp) |
| Newman 2014, (50) | CAPP-Seq | 0.02% | 85% | 96% | Noninvasive tumor genotyping and MRD detection | Tumor-naive assays | 139 genes (125 kb) |
| Newman 2016, (51) | iDES-enhanced CAPP-Seq | 0.004% | 90% | 96% | Noninvasive tumor genotyping and MRD detection | Tumor-naive assays | 128 genes (188 kbp) |
| Phallen 2017, (52) | TEC-Seq | – | 62% | >99% | Detection of early-stage cancers | Tumor-informed assays | 58 genes (80.9 kbp) |
| Zviran 2020, (53) | MRDetect | 0.001% | 67% | 96% | Detection of post-operative MRD | Tumor-informed assays | >10,000 SNVs (>4 Mbp) |
| Kurtz 2021, (55) | PhasED-Seq | 0.000094% | 75% | 97% | Enhanced detection of MRD | Tumor-informed assays | 120 bp hybrid capture panels |
| Chen 2023, (57) | PROPHET | 0.001% | 84.2% | 92.3% | Head-to-head comparison Fixed-panel assays and personalized assays |
Personalized tumor-informed assays | 50 highly ranked variants |
LOD, limit of detection; MRD, minimal residual disease; SNV, single nucleotide variant.
Studies have demonstrated that integrating ctDNA with a panel of biomarkers, including proteins and exosomal RNAs, can significantly enhance detection sensitivity (59,60). For example, The CancerSEEK early cancer detection tool (4), which integrates ctDNA and protein biomarkers, increases the sensitivity of ctDNA-conjugated protein biomarker assays in patients with NSCLC compared to ctDNA analysis alone. Similarly, ctDNA combined with exosomal RNA improved the sensitivity of EGFR mutation detection in patients with advanced NSCLC compared with ctDNA analysis alone (60). In recent years, ctDNA methylation analysis has emerged as a promising tool with significant clinical potential and value. Galleri, developed by Grail, is a widely recognized ctDNA methylation detection platform capable of identifying over 50 different types of cancers across various stages. It demonstrates a detection sensitivity of 43.9% for stage I-III cancers and a specificity of up to 99.3%. This technology has also yielded excellent results in early cancer screening (61,62).
Notably, in patients with brain metastases, the blood-brain barrier prevents the release of ctDNA from central nervous system tumors into the peripheral blood, resulting in a lower detection rate of plasma ctDNA (63). Therefore, overcoming the blood-brain barrier for ctDNA detection is worth further exploration. ctDNA detection based on cerebrospinal fluid (CSF) may be more sensitive than that based on plasma ctDNA (64,65). CSF ctDNA binding to exosomal RNA and T-cell receptors, and simultaneous longitudinal monitoring of the disease response may be a direction for future research.
Biological mutations detected in the plasma might not be tumor cell-derived mutations but rather originated from hematopoietic lineage cells (66). Hematopoietic stem/progenitor cells undergo somatic mutations in their genetic material during division and are released into the blood, which increases with age, particularly in individuals aged >70 years, with mutation rates of up to 5–6% (67,68). Thus, clonal hematopoiesis of indeterminate potential (CHIP) is characterized by the detection of somatic mutations in myeloid-related genes in peripheral blood or bone marrow, with a VAF of at least 2%, but without fulfilling the diagnostic criteria for hematological malignancies as defined by the World Health Organization (WHO) (69). It significantly impacts detection specificity. Studies have demonstrated that clonal hematopoiesis is detected in the plasma of approximately 75% of NSCLC patients (70,71). In a prospective study by Razavi et al., 508 gene combinations for deep sequencing (>60,000× raw-depth) of somatic mutations in 124 patients with metastatic cancer revealed that 53.2% and 81.6% of cfDNA in cancer and non-cancer patients, respectively, originated from hematopoietic lineage cell mutations (72). These mutations involved TP53, JAK2, and KRAS mutations (73). Therefore, one of the main challenges is effective detection of CHIP. The most direct and feasible method to identify and remove CHIP-related mutations and reduce the false-positive rate of ctDNA is deep sequencing of PBMCs.
Detection time
At what time point post-surgery should ctDNA testing be conducted? From a clinical perspective, determining MRD status at an early post-surgery landmark time was highly attractive because it would facilitate immediate decision-making regarding adjuvant therapies (56). Given that surgical trauma inevitably causes systemic inflammatory reactions and damages to the surrounding tissues, leading to an increase in ctDNA concentration. It is difficult to determine whether postoperative ctDNA positivity is MRD or ctDNA has not been completely cleared after surgery (74-76). The DYNAMIC study conducted continuous sampling at multiple time points, including 1 day, 3 days, and 1 month after surgery, and analyzed the ability of postoperative 3-day MRD status to predict the risk of tumor recurrence (77). The MEDAL study showed that compared with 3-day postoperative MRD positivity, 1-month postoperative MRD positivity might be associated with a higher risk of recurrence, and 1-month postoperative MRD status was added to TNM staging to establish tumor-node-metastasis-blood (TNMB) staging that can more accurately stratify the prognosis of NSCLC (P<0.001) (55). A meta-analysis comprehensively collated and analyzed the existing literature data and, based on the analysis results, divided the postoperative detection time points into three stages: 3 days to 2 weeks postoperatively, 1 to 3 months, and every 3 to 6 months. The results showed that in the above three stages, the recurrence risk of MRD+ patients increased to 8.4 times, 6.28 times, and 4.28 times, respectively (78). Therefore, based on the existing literature, it is recommend conducting the first MRD test within the period from the second week to one month after the operation (57,78). Moreover, longitudinal MRD testing at multiple time points during regular follow-up has the advantage of decreasing false negatives probably due to the expansion of potential residual tumor cells during the follow-up. Most published studies have demonstrated that longitudinal MRD testing outperformed single-time-point (landmark) MRD assessments in terms of positive predictive value (PPV) or negative predictive value (NPV) (57,77). As MRD usually can be detectable with a significant lead time over conventional imaging, it would be possible to initiate adjuvant or consolidation therapy at the first positive blood draw during the longitudinal MRD monitoring, and timely intervention can potentially improve patient outcomes by addressing residual disease before it becomes clinically apparent (Table 3).
Table 3
| Studies | Stage | Treatment methods |
N | Lead time (days) | Platform | Sensitivity | Specificity | Trial registration | Panel |
|---|---|---|---|---|---|---|---|---|---|
| Chaudhuri 2017, (25) | IB–III NSCLC | Surgery ± CRT | 40 | 156 | CAPP-Seq | 94% | 100% | NCT01385722 NCT00349830 |
128 genes (188 kbp) |
| Abbosh 2017, (26) | IA–IIIB NSCLC | Surgery ± CRT | 24 | 70 | Signatera | 93% | 90% | NCT01888601 | Median of 18 SNVs |
| Ohara 2020, (28) | IIA–IIIA LC | Surgery ± ICI | 20 | – | CAPP-Seq | 75% | 88% | – | 197 genes |
| Kuang 2020, (29) | IB–III NSCLC | Surgery ± chemo | 36 | – | NGS | 44% | 81% | NCT03465241 | 425 genes |
| Isaksson 2019, (30) | I–IIIA NSCLC | Surgery ± CRT | 58 | – | ddPCR | 80% | 63% | – | 26 genes |
| Xia 2022, (31) | I–III NSCLC | Surgery ± ADT | 330 | – | NGS | 81% | 84% | NCT03317080 | 769 genes |
| Peng 2020, (33) | I–IV LC | Surgery ± chemo/targeted therapy | 77 | 378 | cSMART | 63% | 61% | – | 127 genes |
| Li 2022, (34) | I–IIIA NSCLC | Surgery ± chemo | 119 | 261 | NGS | 64% | 78% | NCT03465241 | 425 genes |
| Zhang 2023, (36) | IA–IIIB NSCLC | Surgery ± chemo | 73 | – | NGS | 71% | 88% | – | – |
| Qiu 2021, (37) | II–III NSCLC | Surgery ± chemo | 103 | 88 | NGS | 79.4% | 89.8% | ChiCTR19000–24656 | 139 genes |
| Zhang 2022, (38) | I–IIIA NSCLC | Surgery ± chemo/targeted therapy | 61 | 102 | NGS | 87.2% (longitudinal) | 97% (longitudinal) | – | 338 genes |
| Bossé 2024, (39) | IA–IB | Surgery | 260 | – | MCED | 36% | 99.4% | – | 1166720 CpGs (17.2 Mb) |
| Moding 2020, (45) | IIB–III NSCLC | CRT ± ICI | 65 | 123 | CAPP-Seq | 82% | 87% | NCT00349830, MDACC LAB09-0983, NCT02525757 | 128 genes |
| Pan 2023, (46) | IIB–IIIC NSCLC | CRT ± ICI/TKI | 139 | 120 | ER-seq | 97.8% (longitudinal) | 53.1% (longitudinal) | NCT04841811 | 1,021 genes |
| Chen 2023, (57) | I–III NSCLC |
Surgery ± ADT | 81 | 299 | PROPHET | 84.2% (longitudinal) | >92.3% (longitudinal) | NCT03634826 | 50 highly ranked variants |
| Chen 2019, (77) | I–IIIA NSCLC | Surgery ± CRT/targeted therapy | 205 | 165 | cSMART | 44% | 88% | NCT02965391 | 8 genes |
ADT, adjuvant therapy; CRT, chemoradiotherapy; ICI, immune checkpoint inhibitor; LC, lung cancer; MRD, minimal residual disease; NSCLC, non-small cell lung cancer; NGS, next-generation sequencing; SNV, single nucleotide variant; TKI, tyrosine kinase inhibitors.
Cut-off of ctDNA MRD assays
For mutation-based ctDNA MRD assays, establishing a cut-off for ctDNA levels was influenced by both biological and technical factors. These included the amount of cfDNA input, sample quality, and the specific experiment or analysis approaches employed. Consequently, it was challenging to establish or compare a common cut-off across different studies. The variability in these factors could lead to differences in sensitivity and specificity, making standardization difficult. Moreover, in scenarios involving treatment escalation or de-escalation, the requirements for predictive values differed significantly (79). Treatment escalation generally demanded a high PPV to ensure that patients identified as MRD-positive could truly benefit from additional therapy. Conversely, treatment de-escalation required a high NPV to ensure that patients classified as MRD-negative were safe to undergo reduced treatment intensity without compromising their outcomes. Balancing these requirements was crucial for optimizing patient care and ensuring that therapeutic decisions were both effective and safe.
In summary, while mutation-based ctDNA MRD assays offered promising tools for personalized medicine, the variability in cut-off determination and the differing needs for PPV and NPV in treatment adjustment highlighted the need for careful consideration and tailored approaches in clinical application.
Feasibility assessment
Given the substantial clinical application potential of ctDNA MRD, ctDNA MRD is likely to become a standard tool for monitoring tumor recurrence and treatment response in the future. Consequently, it is imperative to address an important yet often overlooked challenge: how to integrate ctDNA MRD testing into routine clinical workflows. Firstly, it is crucial to determine whether to outsource these tests or conduct them in-house. Existing studies have performed head-to-head comparisons of these two approaches, revealing that while they exhibit a high degree of consistency, certain mutation discrepancies remain. These differences may ultimately influence clinical decision-making (80,81). These mutation discrepancies may be attributed to various technical factors, including research design, sample quality, enrichment and analysis methodologies, and bioinformatics filtering thresholds (82). In the future, the development of standardized methods with enhanced sensitivity and specificity will present significant opportunities to improve detection rates and clinical applications. Another crucial aspect of incorporating ctDNA MRD into clinical workflows is the education and training of healthcare professionals, including pathologists, physicians, and nurses, to ensure they can accurately apply and interpret the testing methods. Additionally, clinicians who receive the test reports must be trained in reading and interpreting the results and taking appropriate clinical actions. Such training for professionals can be facilitated through specialized workshops held in the workplace or integrated into university curricula as part of ongoing education on emerging technologies. Finally, ctDNA testing involves a complex technical workflow and advanced equipment, encompassing multiple stages such as sample collection, processing, sequencing analysis, and professional interpretation. This intricate process contributes to relatively high testing costs. Consequently, the substantial financial burden associated with ctDNA testing can be significant for patients. To promote the wide application of ctDNA testing, it is crucial to reduce the testing cost and improve the reimbursement policy. Relevant enterprises should lower the cost through technological innovation and process optimization, while the government and medical insurance departments should comprehensively evaluate its clinical value and the economic burden on patients, and gradually incorporate it into the medical insurance reimbursement system to benefit more patients.
Conclusions
Currently, ctDNA MRD provides strong support for recurrence monitoring and efficacy assessment in NSCLC diagnosis and treatment. In terms of monitoring recurrence, it can identify tumor recurrence earlier than imaging. On the other hand, ctDNA MRD testing can be performed to assist in the diagnosis, and a negative MRD indicates a small possibility of recurrence. Moreover, ctDNA MRD can screen patients who need treatment and avoid overtreatment or undertreatment, thereby reducing patients’ adverse reactions to treatment, improving quality of life, and reducing patients’ economic burden.
In order to be applied into clinical practice, ctDNA MRD should solve at least partly the following issues: first, the current technology has low sensitivity to detect low-frequency mutations and early tumors, and the detection cost is expensive. Thus, building an ideal technology with high sensitivity, easy promotion, and low cost is the direction for future research. Second, the timing of postoperative MRD testing varied among patients with lung cancer. We recommend that patients undergoing surgical treatment receive the first MRD test within 2 weeks to 1 month after surgery, and then longitudinal monitoring every 3–6 months; patients with high risk of recurrence after surgery are advised to synchronize the MRD detection time point with the clinical review time point; and locally advanced and advanced patients receiving internal medicine treatment are advised to synchronize the MRD detection time point with the clinical efficacy evaluation time point. Finally, future studies should focus on patient experiences and explore how ctDNA MRD monitoring affects patients’ quality of life, anxiety levels, and treatment adherence, which would be invaluable in supporting the clinical application of ctDNA MRD.
Acknowledgments
None.
Footnote
Reporting Checklist: The authors have completed the Narrative Review reporting checklist. Available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-24-942/rc
Peer Review File: Available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-24-942/prf
Funding: This work was support by
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://tlcr.amegroups.com/article/view/10.21037/tlcr-24-942/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Xia C, Dong X, Li H, et al. Cancer statistics in China and United States, 2022: profiles, trends, and determinants. Chin Med J (Engl) 2022;135:584-90. [Crossref] [PubMed]
- Reck M, Rabe KF. Precision Diagnosis and Treatment for Advanced Non-Small-Cell Lung Cancer. N Engl J Med 2017;377:849-61. [Crossref] [PubMed]
- Zhu S, Wu R, Liu X, et al. Clinical application of ctDNA in early diagnosis, treatment and prognosis of patients with non-small cell lung cancer. Future Oncol 2024;20:2213-24. [Crossref] [PubMed]
- Cohen JD, Li L, Wang Y, et al. Detection and localization of surgically resectable cancers with a multi-analyte blood test. Science 2018;359:926-30. [Crossref] [PubMed]
- Gentzler RD, Guittar J, Mitra A, et al. Dynamic Changes in Circulating Tumor Fraction as a Predictor of Real-World Clinical Outcomes in Solid Tumor Malignancy Patients Treated with Immunotherapy. Oncol Ther 2024;12:509-24. [Crossref] [PubMed]
- Li RY, Liang ZY. Circulating tumor DNA in lung cancer: real-time monitoring of disease evolution and treatment response. Chin Med J (Engl) 2020;133:2476-85. [Crossref] [PubMed]
- Repetto O, Vettori R, Steffan A, et al. Circulating Proteins as Diagnostic Markers in Gastric Cancer. Int J Mol Sci 2023;24:16931. [Crossref] [PubMed]
- Huang K, Dahele M, Senan S, et al. Radiographic Changes After Lung Stereotactic Ablative Radiotherapy (SABR) -- Can We Distinguish Fibrosis From Recurrence? A Systematic Review of the Literature. Pract Radiat Oncol 2013;3:S11-2. [Crossref] [PubMed]
- Long NM, Smith CS. Causes and imaging features of false positives and false negatives on F-PET/CT in oncologic imaging. Insights Imaging 2011;2:679-98. [Crossref] [PubMed]
- Verzè M, Pluchino M, Leonetti A, et al. Role of ctDNA for the detection of minimal residual disease in resected non-small cell lung cancer: a systematic review. Transl Lung Cancer Res 2022;11:2588-600. [Crossref] [PubMed]
- Marinello A, Tagliamento M, Pagliaro A, et al. Circulating tumor DNA to guide diagnosis and treatment of localized and locally advanced non-small cell lung cancer. Cancer Treat Rev 2024;129:102791. [Crossref] [PubMed]
- Ho HY, Chung KK, Kan CM, et al. Liquid Biopsy in the Clinical Management of Cancers. Int J Mol Sci 2024;25:8594. [Crossref] [PubMed]
- Döhner H, Wei AH, Appelbaum FR, et al. Diagnosis and management of AML in adults: 2022 recommendations from an international expert panel on behalf of the ELN. Blood 2022;140:1345-77. [Crossref] [PubMed]
- Kumar S, Paiva B, Anderson KC, et al. International Myeloma Working Group consensus criteria for response and minimal residual disease assessment in multiple myeloma. Lancet Oncol 2016;17:e328-46. [Crossref] [PubMed]
- Tomuleasa C, Selicean C, Cismas S, et al. Minimal residual disease in chronic lymphocytic leukemia: A consensus paper that presents the clinical impact of the presently available laboratory approaches. Crit Rev Clin Lab Sci 2018;55:329-45. [Crossref] [PubMed]
- Hourigan CS, Karp JE. Minimal residual disease in acute myeloid leukaemia. Nat Rev Clin Oncol 2013;10:460-71. [Crossref] [PubMed]
- Adler S, Seidel J, Choyke P, et al. Minimum lesion detectability as a measure of PET system performance. EJNMMI Phys 2017;4:13. [Crossref] [PubMed]
- Yan X, Liu C. Application of Non-Blood-Derived Fluid Biopsy in Monitoring Minimal Residual Diseases of Lung Cancer. Front Surg 2022;9:865040. [Crossref] [PubMed]
- Mouliere F, Chandrananda D, Piskorz AM, et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci Transl Med 2018;10:eaat4921. [Crossref] [PubMed]
- Li C, Shao J, Li P, et al. Circulating tumor DNA as liquid biopsy in lung cancer: Biological characteristics and clinical integration. Cancer Lett 2023;577:216365. [Crossref] [PubMed]
- Sanchez C, Roch B, Mazard T, et al. Circulating nuclear DNA structural features, origins, and complete size profile revealed by fragmentomics. JCI Insight 2021;6:e144561. [Crossref] [PubMed]
- Siravegna G, Marsoni S, Siena S, et al. Integrating liquid biopsies into the management of cancer. Nat Rev Clin Oncol 2017;14:531-48. [Crossref] [PubMed]
- Shen H, Jin Y, Zhao H, et al. Potential clinical utility of liquid biopsy in early-stage non-small cell lung cancer. BMC Med 2022;20:480. [Crossref] [PubMed]
- Pellini B, Chaudhuri AA. Circulating Tumor DNA Minimal Residual Disease Detection of Non-Small-Cell Lung Cancer Treated With Curative Intent. J Clin Oncol 2022;40:567-75. [Crossref] [PubMed]
- Chaudhuri AA, Chabon JJ, Lovejoy AF, et al. Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling. Cancer Discov 2017;7:1394-403. [Crossref] [PubMed]
- Abbosh C, Birkbak NJ, Wilson GA, et al. Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution. Nature 2017;545:446-51. [Crossref] [PubMed]
- Abbosh C, Frankell A, Garnett A, et al. Abstract CT023: Phylogenetic tracking and minimal residual disease detection using ctDNA in early-stage NSCLC: A lung TRACERx study. Can Res 2020;80:CT023.
- Ohara S, Suda K, Sakai K, et al. Prognostic implications of preoperative versus postoperative circulating tumor DNA in surgically resected lung cancer patients: a pilot study. Transl Lung Cancer Res 2020;9:1915-23. [Crossref] [PubMed]
- Kuang PP, Li N, Liu Z, et al. Circulating Tumor DNA Analyses as a Potential Marker of Recurrence and Effectiveness of Adjuvant Chemotherapy for Resected Non-Small-Cell Lung Cancer. Front Oncol 2020;10:595650. [Crossref] [PubMed]
- Isaksson S, George AM, Jönsson M, et al. Pre-operative plasma cell-free circulating tumor DNA and serum protein tumor markers as predictors of lung adenocarcinoma recurrence. Acta Oncol 2019;58:1079-86. [Crossref] [PubMed]
- Xia L, Mei J, Kang R, et al. Perioperative ctDNA-Based Molecular Residual Disease Detection for Non-Small Cell Lung Cancer: A Prospective Multicenter Cohort Study (LUNGCA-1). Clin Cancer Res 2022;28:3308-17. [Crossref] [PubMed]
- Abbosh C, Frankell AM, Harrison T, et al. Tracking early lung cancer metastatic dissemination in TRACERx using ctDNA. Nature 2023;616:553-62. [Crossref] [PubMed]
- Peng M, Huang Q, Yin W, et al. Circulating Tumor DNA as a Prognostic Biomarker in Localized Non-small Cell Lung Cancer. Front Oncol 2020;10:561598. [Crossref] [PubMed]
- Li N, Wang BX, Li J, et al. Perioperative circulating tumor DNA as a potential prognostic marker for operable stage I to IIIA non-small cell lung cancer. Cancer 2022;128:708-18. [Crossref] [PubMed]
- Riely GJ, Wood DE, Ettinger DS, et al. Non-Small Cell Lung Cancer, Version 4.2024, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw 2024;22:249-74. [Crossref] [PubMed]
- Zhang X, Zhang Y, Zhang S, et al. Investigate the application of postoperative ctDNA-based molecular residual disease detection in monitoring tumor recurrence in patients with non-small cell lung cancer--A retrospective study of ctDNA. Front Oncol 2023;13:1098128. [Crossref] [PubMed]
- Qiu B, Guo W, Zhang F, et al. Dynamic recurrence risk and adjuvant chemotherapy benefit prediction by ctDNA in resected NSCLC. Nat Commun 2021;12:6770. [Crossref] [PubMed]
- Zhang JT, Liu SY, Gao W, et al. Longitudinal Undetectable Molecular Residual Disease Defines Potentially Cured Population in Localized Non-Small Cell Lung Cancer. Cancer Discov 2022;12:1690-701. [Crossref] [PubMed]
- Bossé Y, Dasgupta A, Abadier M, et al. Prognostic implication of methylation-based circulating tumor DNA detection prior to surgery in stage I non-small cell lung cancer. Cancer Lett 2024;594:216984. [Crossref] [PubMed]
- Sujit SJ, Aminu M, Karpinets TV, et al. Enhancing NSCLC recurrence prediction with PET/CT habitat imaging, ctDNA, and integrative radiogenomics-blood insights. Nat Commun 2024;15:3152. [Crossref] [PubMed]
- Antonia SJ, Villegas A, Daniel D, et al. Durvalumab after Chemoradiotherapy in Stage III Non-Small-Cell Lung Cancer. N Engl J Med 2017;377:1919-29. [Crossref] [PubMed]
- Spigel DR, Faivre-Finn C, Gray JE, et al. Five-Year Survival Outcomes From the PACIFIC Trial: Durvalumab After Chemoradiotherapy in Stage III Non-Small-Cell Lung Cancer. J Clin Oncol 2022;40:1301-11. [Crossref] [PubMed]
- Soria JC, Ohe Y, Vansteenkiste J, et al. Osimertinib in Untreated EGFR-Mutated Advanced Non-Small-Cell Lung Cancer. N Engl J Med 2018;378:113-25. [Crossref] [PubMed]
- Peters S, Camidge DR, Shaw AT, et al. Alectinib versus Crizotinib in Untreated ALK-Positive Non-Small-Cell Lung Cancer. N Engl J Med 2017;377:829-38. [Crossref] [PubMed]
- Moding EJ, Liu Y, Nabet BY, et al. Circulating Tumor DNA Dynamics Predict Benefit from Consolidation Immunotherapy in Locally Advanced Non-Small Cell Lung Cancer. Nat Cancer 2020;1:176-83. [Crossref] [PubMed]
- Pan Y, Zhang JT, Gao X, et al. Dynamic circulating tumor DNA during chemoradiotherapy predicts clinical outcomes for locally advanced non-small cell lung cancer patients. Cancer Cell 2023;41:1763-1773.e4. [Crossref] [PubMed]
- Dong S, Wang Z, Zhang JT, et al. Circulating Tumor DNA-Guided De-Escalation Targeted Therapy for Advanced Non-Small Cell Lung Cancer: A Nonrandomized Controlled Trial. JAMA Oncol 2024;10:932-40. [Crossref] [PubMed]
- Xu J, Wan R, Cai Y, et al. Circulating tumor DNA-based stratification strategy for chemotherapy plus PD-1 inhibitor in advanced non-small-cell lung cancer. Cancer Cell 2024;42:1598-1613.e4. [Crossref] [PubMed]
- Abbosh C, Birkbak NJ, Swanton C. Early stage NSCLC - challenges to implementing ctDNA-based screening and MRD detection. Nat Rev Clin Oncol 2018;15:577-86. [Crossref] [PubMed]
- Newman AM, Bratman SV, To J, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med 2014;20:548-54. [Crossref] [PubMed]
- Newman AM, Lovejoy AF, Klass DM, et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat Biotechnol 2016;34:547-55. [Crossref] [PubMed]
- Phallen J, Sausen M, Adleff V, et al. Direct detection of early-stage cancers using circulating tumor DNA. Sci Transl Med 2017;9:eaan2415. [Crossref] [PubMed]
- Zviran A, Schulman RC, Shah M, et al. Genome-wide cell-free DNA mutational integration enables ultra-sensitive cancer monitoring. Nat Med 2020;26:1114-24. [Crossref] [PubMed]
- Widman AJ, Shah M, Frydendahl A, et al. Ultrasensitive plasma-based monitoring of tumor burden using machine-learning-guided signal enrichment. Nat Med 2024;30:1655-66. [Crossref] [PubMed]
- Kurtz DM, Soo J. Enhanced detection of minimal residual disease by targeted sequencing of phased variants in circulating tumor DNA. Nat Biotechnol 2021;39:1537-47. [Crossref] [PubMed]
- Moding EJ, Nabet BY, Alizadeh AA, et al. Detecting Liquid Remnants of Solid Tumors: Circulating Tumor DNA Minimal Residual Disease. Cancer Discov 2021;11:2968-86. [Crossref] [PubMed]
- Chen K, Yang F, Shen H, et al. Individualized tumor-informed circulating tumor DNA analysis for postoperative monitoring of non-small cell lung cancer. Cancer Cell 2023;41:1749-1762.e6. [Crossref] [PubMed]
- Gao W, Zhang JT, Cao W, et al. The prognostic value evaluation of a tissue Comprehensive Genomic Profiling (CGP)-informed personalized MRD detection assay in NSCLC. Ann Oncol 2023;34:S736.
- Dudley JC, Diehn M. Detection and Diagnostic Utilization of Cellular and Cell-Free Tumor DNA. Annu Rev Pathol 2021;16:199-222. [Crossref] [PubMed]
- Krug AK, Enderle D, Karlovich C, et al. Improved EGFR mutation detection using combined exosomal RNA and circulating tumor DNA in NSCLC patient plasma. Ann Oncol 2018;29:700-6. [Crossref] [PubMed]
- Liu MC, Oxnard GR, Klein EA, et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann Oncol 2020;31:745-59. [Crossref] [PubMed]
- Schrag D, Beer TM, McDonnell CH 3rd, et al. Blood-based tests for multicancer early detection (PATHFINDER): a prospective cohort study. Lancet 2023;402:1251-60. [Crossref] [PubMed]
- Seyhan AA. Circulating Liquid Biopsy Biomarkers in Glioblastoma: Advances and Challenges. Int J Mol Sci 2024;25:7974. [Crossref] [PubMed]
- Liu X, Mei F, Fang M, et al. Cerebrospinal fluid ctDNA testing shows an advantage over plasma ctDNA testing in advanced non-small cell lung cancer patients with brain metastases. Front Oncol 2023;13:1322635. [Crossref] [PubMed]
- Diaz M, Chudsky S, Pentsova E, et al. Clinical applications of cerebrospinal fluid liquid biopsies in central nervous system tumors. Transl Oncol 2024;41:101881. [Crossref] [PubMed]
- Snyder MW, Kircher M, Hill AJ, et al. Cell-free DNA Comprises an In Vivo Nucleosome Footprint that Informs Its Tissues-Of-Origin. Cell 2016;164:57-68. [Crossref] [PubMed]
- Xie M, Lu C, Wang J, et al. Age-related mutations associated with clonal hematopoietic expansion and malignancies. Nat Med 2014;20:1472-8. [Crossref] [PubMed]
- Steensma DP, Bejar R, Jaiswal S, et al. Clonal hematopoiesis of indeterminate potential and its distinction from myelodysplastic syndromes. Blood 2015;126:9-16. [Crossref] [PubMed]
- Khoury JD, Solary E, Abla O, et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/Dendritic Neoplasms. Leukemia 2022;36:1703-19.
- Croitoru VM, Cazacu IM, Popescu I, et al. Clonal Hematopoiesis and Liquid Biopsy in Gastrointestinal Cancers. Front Med (Lausanne) 2021;8:772166. [Crossref] [PubMed]
- Yaung SJ, Fuhlbrück F, Peterson M, et al. Clonal Hematopoiesis in Late-Stage Non-Small-Cell Lung Cancer and Its Impact on Targeted Panel Next-Generation Sequencing. JCO Precis Oncol 2020;4:1271-9. [Crossref] [PubMed]
- Razavi P, Li BT, Brown DN, et al. High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants. Nat Med 2019;25:1928-37. [Crossref] [PubMed]
- Hu Y, Ulrich BC, Supplee J, et al. False-Positive Plasma Genotyping Due to Clonal Hematopoiesis. Clin Cancer Res 2018;24:4437-43. [Crossref] [PubMed]
- Turabi K, Klute K, Radhakrishnan P. Decoding the Dynamics of Circulating Tumor DNA in Liquid Biopsies. Cancers (Basel) 2024;16:2432. [Crossref] [PubMed]
- Szpechcinski A, Chorostowska-Wynimko J, Kupis W, et al. Quantitative analysis of free-circulating DNA in plasma of patients with resectable NSCLC. Expert Opin Biol Ther 2012;12:S3-9. [Crossref] [PubMed]
- Henriksen TV, Reinert T, Christensen E, et al. The effect of surgical trauma on circulating free DNA levels in cancer patients-implications for studies of circulating tumor DNA. Mol Oncol 2020;14:1670-9. [Crossref] [PubMed]
- Chen K, Zhao H, Shi Y, et al. Perioperative Dynamic Changes in Circulating Tumor DNA in Patients with Lung Cancer (DYNAMIC). Clin Cancer Res 2019;25:7058-67. [Crossref] [PubMed]
- Wang B, Pei J, Wang S, et al. Prognostic potential of circulating tumor DNA detection at different time periods in resectable non-small cell lung cancer: Evidence from a meta-analysis. Crit Rev Oncol Hematol 2022;177:103771. [Crossref] [PubMed]
- Abbosh C, Hodgson D, Doherty GJ, et al. Implementing circulating tumor DNA as a prognostic biomarker in resectable non-small cell lung cancer. Trends Cancer 2024;10:643-54. [Crossref] [PubMed]
- Heeke S, Hofman V, Ilié M, et al. Prospective evaluation of NGS-based liquid biopsy in untreated late stage non-squamous lung carcinoma in a single institution. J Transl Med 2020;18:87. [Crossref] [PubMed]
- Macías M, Cañada-Higueras E, Alegre E, et al. Performance comparison of two next-generation sequencing panels to detect actionable mutations in cell-free DNA in cancer patients. Clin Chem Lab Med 2020;58:1341-8. [Crossref] [PubMed]
- Stetson D, Ahmed A, Xu X, et al. Orthogonal Comparison of Four Plasma NGS Tests With Tumor Suggests Technical Factors are a Major Source of Assay Discordance. JCO Precis Oncol 2019;3:1-9. [Crossref] [PubMed]


